Interferon signature in the blood in inflammatory common variable immune deficiency
- PMID: 24069364
- PMCID: PMC3775732
- DOI: 10.1371/journal.pone.0074893
Interferon signature in the blood in inflammatory common variable immune deficiency
Abstract
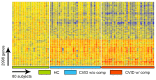
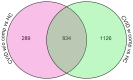
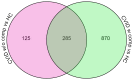
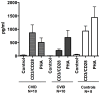
About half of all subjects with common variable immune deficiency (CVID) are afflicted with inflammatory complications including hematologic autoimmunity, granulomatous infiltrations, interstitial lung disease, lymphoid hyperplasia and/or gastrointestinal inflammatory disease. The pathogenesis of these conditions is poorly understood but singly and in aggregate, these lead to significantly increased (11 fold) morbidity and mortality, not experienced by CVID subjects without these complications. To explore the dysregulated networks in these subjects, we applied whole blood transcriptional profiling to 91 CVID subjects, 47 with inflammatory conditions and 44 without, in comparison to subjects with XLA and healthy controls. As compared to other CVID subjects, males with XLA or healthy controls, the signature of CVID subjects with inflammatory complications was distinguished by a marked up-regulation of IFN responsive genes. Chronic up-regulation of IFN pathways is known to occur in autoimmune disease due to activation of TLRs and other still unclarified cytoplasmic sensors. As subjects with inflammatory complications were also more likely to be lymphopenic, have reduced B cell numbers, and a greater reduction of B, T and plasma cell networks, we suggest that more impaired adaptive immunity in these subjects may lead to chronic activation of innate IFN pathways in response to environmental antigens. The unbiased use of whole blood transcriptome analysis may provides a tool for distinguishing CVID subjects who are at risk for increased morbidity and earlier mortality. As more effective therapeutic options are developed, whole blood transcriptome analyses could also provide an efficient means of monitoring the effects of treatment of the inflammatory phenotype.
Conflict of interest statement
Figures






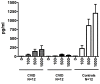
References
-
- Cunningham-Rundles C (2001) Common variable immunodeficiency. Curr Allergy Asthma Rep 1: 421-429. doi:10.1007/s11882-001-0027-1. PubMed: 11892068. - DOI - PubMed
-
- Park JH, Resnick ES, Cunningham-Rundles C (2011) Perspectives on common variable immune deficiency. Ann N Y Acad Sci 1246: 41-49. doi:10.1111/j.1749-6632.2011.06338.x. PubMed: 22236429. - DOI - PMC - PubMed
-
- Grimbacher B, Hutloff A, Schlesier M, Glocker E, Warnatz K et al. (2003) Homozygous loss of ICOS is associated with adult-onset common variable immunodeficiency. Nat Immunol 4: 261-268. doi:10.1038/ni902. PubMed: 12577056. - DOI - PubMed
-
- van Zelm MC, Reisli I, van der Burg M, Castaño D, van Noesel CJ et al. (2006) An antibody-deficiency syndrome due to mutations in the CD19 gene. N Engl J Med 354: 1901-1912. doi:10.1056/NEJMoa051568. PubMed: 16672701. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

