Comprehensive mutation analysis of the CYP21A2 gene: an efficient multistep approach to the molecular diagnosis of congenital adrenal hyperplasia
- PMID: 24071710
- PMCID: PMC5803549
- DOI: 10.1016/j.jmoldx.2013.06.001
Comprehensive mutation analysis of the CYP21A2 gene: an efficient multistep approach to the molecular diagnosis of congenital adrenal hyperplasia
Abstract
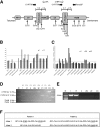
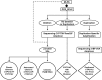
Congenital adrenal hyperplasia, due to 21-hydroxylase deficiency (21-OHD) is an autosomal recessive disorder of adrenal steroidogenesis caused by mutations in the CYP21A2 gene. Direct comparison of established and novel methodologies of CYP21A2 genetic analysis in a large cohort representing a wide range of genotypes has not been previously reported. We genotyped a cohort of 129 unrelated patients with 21-OHD, along with 145 available parents, using Southern blot (SB) analysis, multiplex ligation-dependent probe amplification (MLPA), PCR-based restriction fragment length polymorphism (RFLP) analysis, multiplex minisequencing and conversion-specific PCR, duplication-specific amplification, and DNA sequencing. CYP21A2 genotyping identified four duplicated CYP21A2 genes (1.53%) and 79 chimeric CYP21A1P/CYP21A2 genes (30.15%). Parental SB data were essential for determining the CYP21 haplotype in three cases, whereas PCR-based RFLP analysis was necessary for MLPA results to be accurately interpreted in the majority of cases. The comparison of different methods in detecting deletion and duplication showed that MLPA with PCR-based RFLP was comparable with SB analysis, with parental data of 100% sensitivity and specificity. DNA sequencing was required for the identification of 16 (6.1%) rare point mutations and determination of clinically significant chimera junction sites. MLPA with PCR-based RFLP analysis is an excellent substitute for SB analysis in detecting CYP21A2 deletion and duplication and a combination of MLPA, PCR-based RFLP, duplication-specific amplification, and DNA sequencing is a convenient and comprehensive strategy for mutation analysis of the CYP21A2 gene in patients with 21-OHD.
Published by Elsevier Inc.
Figures

Similar articles
-
A rational, non-radioactive strategy for the molecular diagnosis of congenital adrenal hyperplasia due to 21-hydroxylase deficiency.Gene. 2013 Sep 10;526(2):239-45. doi: 10.1016/j.gene.2013.03.082. Epub 2013 Apr 6. Gene. 2013. PMID: 23570880
-
CYP21A2 mutation analysis in Korean patients with congenital adrenal hyperplasia using complementary methods: sequencing after long-range PCR and restriction fragment length polymorphism analysis with multiple ligation-dependent probe amplification assay.Ann Lab Med. 2015 Sep;35(5):535-9. doi: 10.3343/alm.2015.35.5.535. Ann Lab Med. 2015. PMID: 26206692 Free PMC article.
-
Junction site analysis of chimeric CYP21A1P/CYP21A2 genes in 21-hydroxylase deficiency.Clin Chem. 2012 Feb;58(2):421-30. doi: 10.1373/clinchem.2011.174037. Epub 2011 Dec 7. Clin Chem. 2012. PMID: 22156666 Free PMC article.
-
Variants of the CYP21A2 and CYP21A1P genes in congenital adrenal hyperplasia.Clin Chim Acta. 2013 Mar 15;418:37-44. doi: 10.1016/j.cca.2012.12.030. Epub 2013 Jan 9. Clin Chim Acta. 2013. PMID: 23313747 Review.
-
Molecular diagnosis of congenital adrenal hyperplasia due to 21-hydroxylase deficiency: an update of new CYP21A2 mutations.Clin Chem Lab Med. 2010 Aug;48(8):1057-62. doi: 10.1515/CCLM.2010.239. Clin Chem Lab Med. 2010. PMID: 20482300 Review.
Cited by
-
Mutation detection of CYP21A2 gene in nonclassical congenital adrenal hyperplasia patients with premature pubarche.Adv Biomed Res. 2016 Mar 16;5:33. doi: 10.4103/2277-9175.178794. eCollection 2016. Adv Biomed Res. 2016. PMID: 27099846 Free PMC article.
-
Health problems of adolescent and adult patients with 21-hydroxylase deficiency.Clin Pediatr Endocrinol. 2018;27(4):203-213. doi: 10.1297/cpe.27.203. Epub 2018 Oct 30. Clin Pediatr Endocrinol. 2018. PMID: 30393437 Free PMC article.
-
Congenital adrenal hyperplasia due to two rare CYP21A2 variant alleles, including a novel attenuated CYP21A1P/CYP21A2 chimera.Mol Genet Genomic Med. 2023 Jul;11(7):e2195. doi: 10.1002/mgg3.2195. Epub 2023 May 8. Mol Genet Genomic Med. 2023. PMID: 37157918 Free PMC article.
-
Congenital Adrenal Hyperplasia.J Pediatr Adolesc Gynecol. 2017 Oct;30(5):520-534. doi: 10.1016/j.jpag.2017.04.001. Epub 2017 Apr 24. J Pediatr Adolesc Gynecol. 2017. PMID: 28450075 Free PMC article. Review.
-
Challenges of CYP21A2 genotyping in children with 21-hydroxylase deficiency: determination of genotype-phenotype correlation using next generation sequencing in Southeastern Anatolia.J Endocrinol Invest. 2021 Nov;44(11):2395-2405. doi: 10.1007/s40618-021-01546-z. Epub 2021 Mar 6. J Endocrinol Invest. 2021. PMID: 33677812
References
-
- Merke D.P., Bornstein S.R. Congenital adrenal hyperplasia. Lancet. 2005;365:2125–2136. - PubMed
-
- White P.C., Speiser P.W. Congenital adrenal hyperplasia due to 21-hydroxylase deficiency. Endocr Rev. 2000;21:245–291. - PubMed
-
- Speiser P.W., Azziz R., Baskin L.S., Ghizzoni L., Hensle T.W., Merke D.P., Meyer-Bahlburg H.F., Miller W.L., Montori V.M., Oberfield S.E., Ritzen M., White P.C. Congenital adrenal hyperplasia due to steroid 21-hydroxylase deficiency: an Endocrine Society clinical practice guideline. J Clin Endocrinol Metab. 2010;95:4133–4160. - PMC - PubMed
-
- Yang Z., Mendoza A.R., Welch T.R., Zipf W.B., Yu C.Y. Modular variations of the human major histocompatibility complex class III genes for serine/threonine kinase RP, complement component C4, steroid 21-hydroxylase CYP21, and tenascin TNX (the RCCX module). A mechanism for gene deletions and disease associations. J Biol Chem. 1999;274:12147–12156. - PubMed
-
- White P.C., Tusie-Luna M.T., New M.I., Speiser P.W. Mutations in steroid 21-hydroxylase (CYP21) Hum Mutat. 1994;3:373–378. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

