Nodulin 22, a novel small heat-shock protein of the endoplasmic reticulum, is linked to the unfolded protein response in common bean
- PMID: 24073881
- PMCID: PMC4028047
- DOI: 10.1094/MPMI-07-13-0200-R
Nodulin 22, a novel small heat-shock protein of the endoplasmic reticulum, is linked to the unfolded protein response in common bean
Abstract
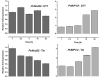
The importance of plant small heat shock proteins (sHsp) in multiple cellular processes has been evidenced by their unusual abundance and diversity; however, little is known about their biological role. Here, we characterized the in vitro chaperone activity and subcellular localization of nodulin 22 of Phaseolus vulgaris (PvNod22; common bean) and explored its cellular function through a virus-induced gene silencing-based reverse genetics approach. We established that PvNod22 facilitated the refolding of a model substrate in vitro, suggesting that it acts as a molecular chaperone in the cell. Through microscopy analyses of PvNod22, we determined its localization in the endoplasmic reticulum (ER). Furthermore, we found that silencing of PvNod22 resulted in necrotic lesions in the aerial organs of P. vulgaris plants cultivated under optimal conditions and that downregulation of PvNod22 activated the ER-unfolded protein response (UPR) and cell death. We also established that PvNod22 expression in wild-type bean plants was modulated by abiotic stress but not by chemicals that trigger the UPR, indicating PvNod22 is not under UPR control. Our results suggest that the ability of PvNod22 to suppress protein aggregation contributes to the maintenance of ER homeostasis, thus preventing the induction of cell death via UPR in response to oxidative stress during plant-microbe interactions.
Figures

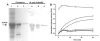




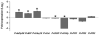
References
-
- Borges A, Tsai SM, Caldas DG. Validation of reference genes for RTqPCR normalization in common bean during biotic and abiotic stresses. Plant Cell Rep. 2012;31:827–838. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

