Tolerance of Escherichia coli to fluoroquinolone antibiotics depends on specific components of the SOS response pathway
- PMID: 24077306
- PMCID: PMC3832272
- DOI: 10.1534/genetics.113.152306
Tolerance of Escherichia coli to fluoroquinolone antibiotics depends on specific components of the SOS response pathway
Abstract
Bacteria exposed to bactericidal fluoroquinolone (FQ) antibiotics can survive without becoming genetically resistant. Survival of these phenotypically resistant cells, commonly called "persisters," depends on the SOS gene network. We have examined mutants in all known SOS-regulated genes to identify functions essential for tolerance in Escherichia coli. The absence of DinG and UvrD helicases and the Holliday junction processing enzymes RuvA and RuvB leads to a decrease in survival. Analysis of the respective mutants indicates that, in addition to repair of double-strand breaks, tolerance depends on the repair of collapsed replication forks and stalled transcription complexes. Mutation in recF results in increased survival, which identifies RecAF recombination as a poisoning mechanism not previously linked to FQ lethality. DinG acts upstream of SOS promoting its induction, whereas RuvAB participates in repair only. UvrD directly promotes all repair processes initiated by FQ-induced damage and prevents RecAF-dependent misrepair, making it one of the crucial SOS functions required for tolerance.
Keywords: DNA repair; SOS response; fluoroquinolones; persisters; tolerance.
Figures
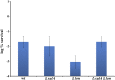
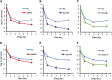

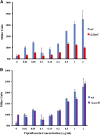
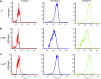

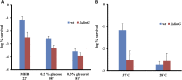

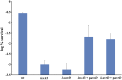
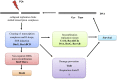
Similar articles
-
Analysis of RuvABC and RecG involvement in the escherichia coli response to the covalent topoisomerase-DNA complex.J Bacteriol. 2010 Sep;192(17):4445-51. doi: 10.1128/JB.00350-10. Epub 2010 Jul 2. J Bacteriol. 2010. PMID: 20601468 Free PMC article.
-
Fluoroquinolone-specific resistance trajectories in E. coli and their dependence on the SOS-response.BMC Microbiol. 2025 Jan 21;25(1):37. doi: 10.1186/s12866-025-03771-5. BMC Microbiol. 2025. PMID: 39838279 Free PMC article.
-
lon incompatibility associated with mutations causing SOS induction: null uvrD alleles induce an SOS response in Escherichia coli.J Bacteriol. 2000 Jun;182(11):3151-7. doi: 10.1128/JB.182.11.3151-3157.2000. J Bacteriol. 2000. PMID: 10809694 Free PMC article.
-
Biology before the SOS Response-DNA Damage Mechanisms at Chromosome Fragile Sites.Cells. 2021 Sep 1;10(9):2275. doi: 10.3390/cells10092275. Cells. 2021. PMID: 34571923 Free PMC article. Review.
-
Properties of the Escherichia coli RuvA and RuvB proteins involved in DNA repair, recombination and mutagenesis.Biochimie. 1991 Apr;73(4):505-7. doi: 10.1016/0300-9084(91)90120-p. Biochimie. 1991. PMID: 1911951 Review.
Cited by
-
A 5+1 assemble-to-activate mechanism of the Lon proteolytic machine.Nat Commun. 2023 Nov 13;14(1):7340. doi: 10.1038/s41467-023-43035-2. Nat Commun. 2023. PMID: 37957149 Free PMC article.
-
Cellular Self-Digestion and Persistence in Bacteria.Microorganisms. 2021 Oct 31;9(11):2269. doi: 10.3390/microorganisms9112269. Microorganisms. 2021. PMID: 34835393 Free PMC article. Review.
-
Discovery of antibiotics that selectively kill metabolically dormant bacteria.Cell Chem Biol. 2024 Apr 18;31(4):712-728.e9. doi: 10.1016/j.chembiol.2023.10.026. Epub 2023 Nov 28. Cell Chem Biol. 2024. PMID: 38029756 Free PMC article.
-
Fluoroquinolone Persistence in Escherichia coli Requires DNA Repair despite Differing between Starving Populations.Microorganisms. 2022 Jan 26;10(2):286. doi: 10.3390/microorganisms10020286. Microorganisms. 2022. PMID: 35208744 Free PMC article.
-
Formation of Escherichia coli O157:H7 Persister Cells in the Lettuce Phyllosphere and Application of Differential Equation Models To Predict Their Prevalence on Lettuce Plants in the Field.Appl Environ Microbiol. 2020 Jan 7;86(2):e01602-19. doi: 10.1128/AEM.01602-19. Print 2020 Jan 7. Appl Environ Microbiol. 2020. PMID: 31704677 Free PMC article.
References
-
- Bremer H., Dennis P. P., 1987. Modulation of chemical composition and other parameters of the cell by growth rate, pp. 1527–1542 in Escherichia coli and Salmonella typhimurium: Cellular and Molecular Biology, edited by Neidhardt F. C. American Society for Microbiology, Washington, DC.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

