Human GEN1 and the SLX4-associated nucleases MUS81 and SLX1 are essential for the resolution of replication-induced Holliday junctions
- PMID: 24080495
- PMCID: PMC3844290
- DOI: 10.1016/j.celrep.2013.08.041
Human GEN1 and the SLX4-associated nucleases MUS81 and SLX1 are essential for the resolution of replication-induced Holliday junctions
Abstract
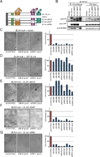
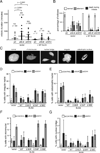
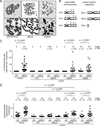
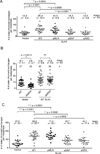
Holliday junctions (HJs), the DNA intermediates of homologous recombination, need to be faithfully processed in order to preserve genome integrity. In human cells, the BLM helicase complex promotes nonnucleolytic dissolution of double HJs. In vitro, HJs may be nucleolytically processed by MUS81-EME1, GEN1, and SLX4-SLX1. Here, we exploit human SLX4-null cells to examine the requirements for HJ resolution in vivo. Lack of BLM and SLX4 or GEN1 and SLX4 is synthetically lethal in the absence of exogenous DNA damage, and lethality is a consequence of dysfunctional mitosis proceeding in the presence of unprocessed HJs. Thus, GEN1 activity cannot be substituted for the SLX4-associated nucleases, and one of the HJ resolvase activities, either of those associated with SLX4 or with GEN1, is required for cell viability, even in the presence of BLM. In vivo HJ resolution depends on both SLX4-associated MUS81-EME1 and SLX1, suggesting that they are acting in concert in the context of SLX4.
Copyright © 2013 The Authors. Published by Elsevier Inc. All rights reserved.
Figures

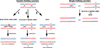
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

