Mechanism of amyloid β-protein dimerization determined using single-molecule AFM force spectroscopy
- PMID: 24096987
- PMCID: PMC3791449
- DOI: 10.1038/srep02880
Mechanism of amyloid β-protein dimerization determined using single-molecule AFM force spectroscopy
Abstract
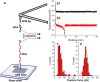
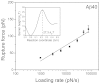
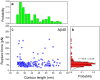
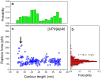
Aβ42 and Aβ40 are the two primary alloforms of human amyloid β-protein (Aβ). The two additional C-terminal residues of Aβ42 result in elevated neurotoxicity compared with Aβ40, but the molecular mechanism underlying this effect remains unclear. Here, we used single-molecule force microscopy to characterize interpeptide interactions for Aβ42 and Aβ40 and corresponding mutants. We discovered a dramatic difference in the interaction patterns of Aβ42 and Aβ40 monomers within dimers. Although the sequence difference between the two peptides is at the C-termini, the N-terminal segment plays a key role in the peptide interaction in the dimers. This is an unexpected finding as N-terminal was considered as disordered segment with no effect on the Aβ peptide aggregation. These novel properties of Aβ proteins suggests that the stabilization of N-terminal interactions is a switch in redirecting of amyloids form the neurotoxic aggregation pathway, opening a novel avenue for the disease preventions and treatments.
Figures
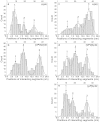
References
-
- Chiti F. & Dobson C. M. Protein misfolding, functional amyloid, and human disease. Annu Rev Biochem 75, 333–366 (2006). - PubMed
-
- Benilova I., Karran E. & De Strooper B. The toxic Abeta oligomer and Alzheimer's disease: an emperor in need of clothes. Nat Neurosci 15, 349–357 (2012). - PubMed
-
- Karran E., Mercken M. & De Strooper B. The amyloid cascade hypothesis for Alzheimer's disease: an appraisal for the development of therapeutics. Nat Rev Drug Discov 10, 698–712 (2011). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

