In vitro phenotypic characterization of hepatitis C virus NS3 protease variants observed in clinical studies of telaprevir
- PMID: 24100495
- PMCID: PMC3837892
- DOI: 10.1128/AAC.01578-13
In vitro phenotypic characterization of hepatitis C virus NS3 protease variants observed in clinical studies of telaprevir
Abstract
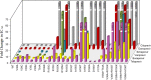
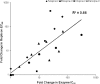
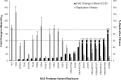
Telaprevir is a linear, peptidomimetic small molecule that inhibits hepatitis C virus (HCV) replication by specifically inhibiting the NS3·4A protease. In phase 3 clinical studies, telaprevir in combination with peginterferon and ribavirin (PR) significantly improved sustained virologic response (SVR) rates in genotype 1 chronic HCV-infected patients compared with PR alone. In patients who do not achieve SVR after treatment with telaprevir-based regimens, variants with mutations in the NS3·4A protease region have been observed. Such variants can contribute to drug resistance and limit the efficacy of treatment. To gain a better understanding of the viral resistance profile, we conducted phenotypic characterization of the variants using HCV replicons carrying site-directed mutations. The most frequently observed (significantly enriched) telaprevir-resistant variants, V36A/M, T54A/S, R155K/T, and A156S, conferred lower-level resistance (3- to 25-fold), whereas A156T and V36M+R155K conferred higher-level resistance (>25-fold) to telaprevir. Rarely observed (not significantly enriched) variants included V36I/L and I132V, which did not confer resistance to telaprevir; V36C/G, R155G/I/M/S, V36A+T54A, V36L+R155K, T54S+R155K, and R155T+D168N, which conferred lower-level resistance to telaprevir; and A156F/N/V, V36A+R155K/T, V36M+R155T, V36A/M+A156T, T54A+A156S, T54S+A156S/T, and V36M+T54S+R155K, which conferred higher-level resistance to telaprevir. All telaprevir-resistant variants remained fully sensitive to alpha interferon, ribavirin, and HCV NS5B nucleoside and nonnucleoside polymerase inhibitors. In general, the replication capacity of telaprevir-resistant variants was lower than that of the wild-type replicon.
Figures
References
-
- Alter HJ, Seeff LB. 2000. Recovery, persistence, and sequelae in hepatitis C virus infection: a perspective on long-term outcome. Semin. Liver Dis. 20:17–35 - PubMed
-
- Ghobrial RM, Steadman R, Gornbein J, Lassman C, Holt CD, Chen P, Farmer DG, Yersiz H, Danino N, Collisson E, Baquarizo A, Han SS, Saab S, Goldstein LI, Donovan JA, Esrason K, Busuttil RW. 2001. A 10-year experience of liver transplantation for hepatitis C: analysis of factors determining outcome in over 500 patients. Ann. Surg. 234:384–393 - PMC - PubMed
-
- Kenny-Walsh E. 2001. The natural history of hepatitis C virus infection. Clin. Liver Dis. 5:969–977 - PubMed
-
- Purcell RH. 1994. Hepatitis C virus: historical perspective and current concepts. FEMS Microbiol. Rev. 14:181–191 - PubMed
-
- Wasley A, Alter MJ. 2000. Epidemiology of hepatitis C: geographic differences and temporal trends. Semin. Liver Dis. 20:1–16 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

