Characterization of molecular and cellular functions of the cyclin-dependent kinase CDK9 using a novel specific inhibitor
- PMID: 24102143
- PMCID: PMC3874696
- DOI: 10.1111/bph.12408
Characterization of molecular and cellular functions of the cyclin-dependent kinase CDK9 using a novel specific inhibitor
Abstract
Background and purpose: The cyclin-dependent kinase CDK9 is an important therapeutic target but currently available inhibitors exhibit low specificity and/or narrow therapeutic windows. Here we have used a new highly specific CDK9 inhibitor, LDC000067 to interrogate gene control mechanisms mediated by CDK9.
Experimental approach: The selectivity of LDC000067 was established in functional kinase assays. Functions of CDK9 in gene expression were assessed with in vitro transcription experiments, single gene analyses and genome-wide expression profiling. Cultures of mouse embryonic stem cells, HeLa cells, several cancer cell lines, along with cells from patients with acute myelogenous leukaemia were also used to investigate cellular responses to LDC000067.
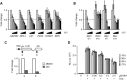
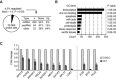
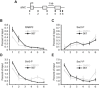
Key results: The selectivity of LDC000067 for CDK9 over other CDKs exceeded that of the known inhibitors flavopiridol and DRB. LDC000067 inhibited in vitro transcription in an ATP-competitive and dose-dependent manner. Gene expression profiling of cells treated with LDC000067 demonstrated a selective reduction of short-lived mRNAs, including important regulators of proliferation and apoptosis. Analysis of de novo RNA synthesis suggested a wide ranging positive role of CDK9. At the molecular and cellular level, LDC000067 reproduced effects characteristic of CDK9 inhibition such as enhanced pausing of RNA polymerase II on genes and, most importantly, induction of apoptosis in cancer cells.
Conclusions and implications: Our study provides a framework for the mechanistic understanding of cellular responses to CDK9 inhibition. LDC000067 represents a promising lead for the development of clinically useful, highly specific CDK9 inhibitors.
Keywords: CDK9; RNA polymerase II; cyclin-dependent kinase; gene expression; transcription.
© 2013 The British Pharmacological Society.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

