Fluorescence spectroscopy of soluble E. coli SPase I Δ2-75 reveals conformational changes in response to ligand binding
- PMID: 24115229
- PMCID: PMC3949126
- DOI: 10.1002/prot.24429
Fluorescence spectroscopy of soluble E. coli SPase I Δ2-75 reveals conformational changes in response to ligand binding
Abstract
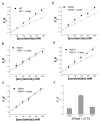
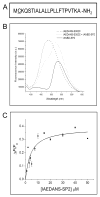
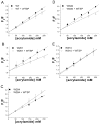
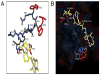
The bacterial Sec pathway is responsible for the translocation of secretory preproteins. During the later stages of transport, the membrane-embedded signal peptidase I (SPase I) cleaves the signal peptide from a preprotein. We used tryptophan fluorescence spectroscopy of a soluble, catalytically active E. coli SPase I Δ2-75 enzyme to study its dynamic conformational changes while in solution and when interacting with lipids and signal peptides. We generated four single Trp SPase I Δ2-75 mutants, W261, W284, W300, and W310. Based on fluorescence quenching experiments, W300 and W310 were found to be more solvent accessible than W261 and W284 in the absence of ligands. W300 and W310 inserted into lipids, consistent with their location at the enzyme's proposed membrane-interface region, while the solvent accessibilities of W261, W284, and W300 were modified in the presence of signal peptide, suggesting propagation of structural changes beyond the active site in response to peptide binding. The signal peptide binding affinity for the enzyme was measured via FRET experiments and the Kd determined to be 4.4 μM. The location of the peptide with respect to the enzyme was also established; this positioning is crucial for the peptide to gain access to the enzyme active site as it emerges from the translocon into the membrane bilayer. These studies reveal enzymatic structural changes required for preprotein proteolysis as it interacts with its two key partners, the signal peptide and membrane phospholipids.
Keywords: acrylamide quenching; preprotein cleavage; protein transport; signal peptidase; signal peptide.
Copyright © 2013 Wiley Periodicals, Inc.
Figures


References
-
- Driessen AJ, Nouwen N. Protein translocation across the bacterial cytoplasmic membrane. Annu Rev Biochem. 2008;77:643–667. - PubMed
-
- Taheri T, Salmanian AH, Gholami E, Doustdari F, Zahedifard F, Rafati S. Leishmania major: disruption of signal peptidase type I and its consequences on survival, growth and infectivity. Exp Parasitol. 2010;126(2):135–145. - PubMed
-
- Paetzel M, Karla A, Strynadka NC, Dalbey RE. Signal peptidases. Chem Rev. 2002;102(12):4549–4580. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

