Structural differences between the avian and human H7N9 hemagglutinin proteins are attributable to modifications in salt bridge formation: a computational study with implications in viral evolution
- PMID: 24116152
- PMCID: PMC3792060
- DOI: 10.1371/journal.pone.0076764
Structural differences between the avian and human H7N9 hemagglutinin proteins are attributable to modifications in salt bridge formation: a computational study with implications in viral evolution
Abstract
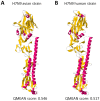
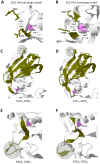
Influenza A hemagglutinin (HA) is a homotrimeric glycoprotein composed of a fibrous globular stem supporting a globular head containing three sialic acid binding sites responsible for infection. The H7N9 strain has consistently infected an avian host, however, the novel 2013 strain is now capable of infecting a human host which would imply that the HA in both strains structurally differ. A better understanding of the structural differences between the avian and human H7N9 strains may shed light into viral evolution and transmissibility. In this study, we elucidated the structural differences between the avian and human H7N9 strains. Throughout the study, we generated HA homology models, verified the quality of each model, superimposed HA homology models to determine structural differences, and, likewise, elucidated the probable cause for these structural differences. We detected two different types of structural differences between the novel H7N9 human and representative avian strains, wherein, one type (Pattern-1) showed three non-overlapping regions while the other type (Pattern-2) showed only one non-overlapping region. In addition, we found that superimposed HA homology models exhibiting Pattern-1 contain three non-overlapping regions designated as: Region-1 (S1571-A1601); Region-3 (R2621-S2651); and Region-4 (S2701-D2811), whereas, superimposed HA homology models showing Pattern-2 only contain one non-overlapping region designated as Region-2 (S1371-S1451). We attributed the two patterns we observed to either the presence of salt bridges involving the E1141 residue or absence of the R1411:D771 salt bridge. Interestingly, comparison between the human H7N7 and H7N9 HA homology models showed high structural similarity. We propose that the putative absence of the R1411:D771 salt bridge coupled with the putative presence of the E1141:R2621 and E1141:K2641 salt bridges found in the 2013 H7N9 HA homology model is associated to human-type receptor binding. This highlights the possible significance of HA salt bridge formation modifications in viral infectivity, immune escape, transmissibility and evolution.
Conflict of interest statement
Figures


Similar articles
-
Effects of HA and NA glycosylation pattern changes on the transmission of avian influenza A(H7N9) virus in guinea pigs.Biochem Biophys Res Commun. 2016 Oct 14;479(2):192-197. doi: 10.1016/j.bbrc.2016.09.024. Epub 2016 Sep 6. Biochem Biophys Res Commun. 2016. PMID: 27613087
-
[Genetic characteristic of hemagglutinin of avian influenza A (H7N9) virus in Guizhou Province in 2017].Zhonghua Yu Fang Yi Xue Za Zhi. 2019 Feb 6;53(2):229-232. doi: 10.3760/cma.j.issn.0253-9624.2019.02.020. Zhonghua Yu Fang Yi Xue Za Zhi. 2019. PMID: 30744302 Chinese.
-
Structures and receptor binding of hemagglutinins from human-infecting H7N9 influenza viruses.Science. 2013 Oct 11;342(6155):243-7. doi: 10.1126/science.1242917. Epub 2013 Sep 5. Science. 2013. PMID: 24009358
-
The Effects of Genetic Variation on H7N9 Avian Influenza Virus Pathogenicity.Viruses. 2020 Oct 28;12(11):1220. doi: 10.3390/v12111220. Viruses. 2020. PMID: 33126529 Free PMC article. Review.
-
Enabling the 'host jump': structural determinants of receptor-binding specificity in influenza A viruses.Nat Rev Microbiol. 2014 Dec;12(12):822-31. doi: 10.1038/nrmicro3362. Epub 2014 Nov 10. Nat Rev Microbiol. 2014. PMID: 25383601 Review.
Cited by
-
Recent Advances in AIV Biosensors Composed of Nanobio Hybrid Material.Micromachines (Basel). 2018 Dec 9;9(12):651. doi: 10.3390/mi9120651. Micromachines (Basel). 2018. PMID: 30544883 Free PMC article. Review.
References
-
- Daniels R, Kurowski B, Johnson AE, Hebert DN (2003) N-linked glycans direct the cotranslational folding pathway of influenza hemagglutinin. Mol Cell 11: 79–90. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

