An upstream YY1 binding site on the HIV-1 LTR contributes to latent infection
- PMID: 24116200
- PMCID: PMC3792934
- DOI: 10.1371/journal.pone.0077052
An upstream YY1 binding site on the HIV-1 LTR contributes to latent infection
Abstract
During HIV-1 infection a population of latently infected cells is established. This population is the major obstacle preventing total eradication of the virus from AIDS patients. HIV-1 latency is thought to arise by various mechanisms including repressive chromatin modifications. Transcription factors such as YY1 have been shown to facilitate repressive chromatin modifications by the recruitment of histone deacetylases. In this study, we identified a novel binding site for YY1 on the HIV-1 LTR, 120 nucleotides upstream of the transcription start site. We show that YY1 can bind to this site in vitro and in vivo and that binding to the LTR is dissociated upon T cell activation. Overexpression of YY1 causes an increase in the proportion of cells that produce latent infections. These observations, in combination with previous results, demonstrate that YY1 plays a prominent role in controlling the establishment and maintenance of latent HIV-1 provirus in unstimulated cells.
Conflict of interest statement
Figures





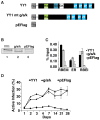

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

