Biallelic genome modification in F(0) Xenopus tropicalis embryos using the CRISPR/Cas system
- PMID: 24123579
- PMCID: PMC4039559
- DOI: 10.1002/dvg.22719
Biallelic genome modification in F(0) Xenopus tropicalis embryos using the CRISPR/Cas system
Abstract
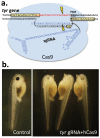
Gene inactivation is an important tool for correlation of phenotypic and genomic data, allowing researchers to infer normal gene function based on the phenotype when the gene is impaired. New and better approaches are needed to overcome the shortfalls of existing methods for any significant acceleration of scientific progress. We have adapted the CRISPR/Cas system for use in Xenopus tropicalis and report on the efficient creation of mutations in the gene encoding the enzyme tyrosinase, which is responsible for oculocutaneous albinism. Biallelic mutation of this gene was detected in the F0 generation, suggesting targeting efficiencies similar to that of TALENs. We also find that off-target mutagenesis seems to be negligible, and therefore, CRISPR/Cas may be a useful system for creating genome modifications in this important model organism.
Keywords: CRISPR/Cas; TALENs; gene knockouts; gene targeting; zinc finger nucleases.
Copyright © 2013 Wiley Periodicals, Inc.
Figures
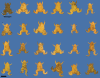

References
-
- Bhaya D, Davison M, Barrangou R. CRISPR-Cas systems in bacteria and archaea: versatile small RNAs for adaptive defense and regulation. Annu Rev Genet. 2011;45:273–97. - PubMed
-
- Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, Lahaye T, Nickstadt A, Bonas U. Breaking the code of DNA binding specificity of TAL-type III effectors. Science. 2009;326:1509–12. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

