Novel small RNA expression libraries uncover hsa-miR-30b and hsa-miR-30c as important factors in anoikis resistance
- PMID: 24129493
- PMCID: PMC3884670
- DOI: 10.1261/rna.039461.113
Novel small RNA expression libraries uncover hsa-miR-30b and hsa-miR-30c as important factors in anoikis resistance
Abstract
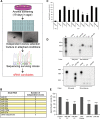
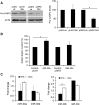
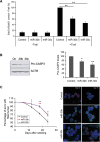
MicroRNAs (miRNAs) have been widely studied in order to elucidate their biological functions. MicroRNA microarrays or miRNA overexpression libraries generated by synthesis and cloning of individual miRNAs have been used to study their different roles. In this work, we have developed a novel methodology to express mature miRNAs and other small RNAs from a double convergent RNA polymerase III promoter. We show that the generated miRNAs function similarly to those processed from primary transcripts or pri-miRNAs. This system allowed us to produce a lentiviral library expressing the whole population of small RNAs present in a metastatic cell line. A functional screening using this library led to the identification of hsa-miR-30b and hsa-miR-30c as negative regulators of cell death induced by loss of attachment (anoikis). Importantly, we demonstrated that the acquisition of anoikis resistance via these miRNAs is achieved through down-regulation of caspase 3 expression. Moreover, overexpression of these miRNAs resulted in a decrease of other types of caspase 3-dependent cell death and enhanced the survival of MCF10A acinar cells in morphogenesis assays, suggesting a putative role as oncomirs. In summary, this novel methodology provides a powerful and effective way for identifying novel small RNAs involved in a particular biological process.
Keywords: anoikis; functional screens; libraries; miR-30b/c; small RNAs.
Figures




Similar articles
-
Large-scale screening identifies a novel microRNA, miR-15a-3p, which induces apoptosis in human cancer cell lines.RNA Biol. 2013 Feb;10(2):287-300. doi: 10.4161/rna.23339. Epub 2013 Jan 25. RNA Biol. 2013. PMID: 23353574 Free PMC article.
-
MicroRNA-141 enhances anoikis resistance in metastatic progression of ovarian cancer through targeting KLF12/Sp1/survivin axis.Mol Cancer. 2017 Jan 17;16(1):11. doi: 10.1186/s12943-017-0582-2. Mol Cancer. 2017. PMID: 28095864 Free PMC article.
-
Chi-miR-30b-5p inhibits dermal papilla cells proliferation by targeting CaMKIIδ gene in cashmere goat.BMC Genomics. 2020 Jun 26;21(1):430. doi: 10.1186/s12864-020-06799-1. BMC Genomics. 2020. PMID: 32586272 Free PMC article.
-
Regulation of the MIR155 host gene in physiological and pathological processes.Gene. 2013 Dec 10;532(1):1-12. doi: 10.1016/j.gene.2012.12.009. Epub 2012 Dec 14. Gene. 2013. PMID: 23246696 Review.
-
Evaluation of MicroRNAs Regulating Anoikis Pathways and Its Therapeutic Potential.Biomed Res Int. 2015;2015:716816. doi: 10.1155/2015/716816. Epub 2015 Oct 26. Biomed Res Int. 2015. PMID: 26587543 Free PMC article. Review.
Cited by
-
Interdigital tissue remodelling in the embryonic limb involves dynamic regulation of the miRNA profiles.J Anat. 2017 Aug;231(2):275-286. doi: 10.1111/joa.12629. Epub 2017 May 24. J Anat. 2017. PMID: 28543398 Free PMC article.
-
Confirmation of a metastasis-specific microRNA signature in primary colon cancer.Sci Rep. 2018 Mar 27;8(1):5242. doi: 10.1038/s41598-018-22532-1. Sci Rep. 2018. PMID: 29588449 Free PMC article.
-
The Expression Patterns of BECN1, LAMP2, and PINK1 Genes in Colorectal Cancer Are Potentially Regulated by Micrornas and CpG Islands: An In Silico Study.J Clin Med. 2020 Dec 12;9(12):4020. doi: 10.3390/jcm9124020. J Clin Med. 2020. PMID: 33322704 Free PMC article.
-
The Role of microRNA Markers in the Diagnosis, Treatment, and Outcome Prediction of Spinal Cord Injury.Front Surg. 2016 Nov 8;3:56. doi: 10.3389/fsurg.2016.00056. eCollection 2016. Front Surg. 2016. PMID: 27878119 Free PMC article. Review.
-
Differential MicroRNA Analyses of Burkholderia pseudomallei- and Francisella tularensis-Exposed hPBMCs Reveal Potential Biomarkers.Int J Genomics. 2017;2017:6489383. doi: 10.1155/2017/6489383. Epub 2017 Jul 16. Int J Genomics. 2017. PMID: 28791299 Free PMC article.
References
-
- Alfano D, Iaccarino I, Stoppelli MP. 2006. Urokinase signaling through its receptor protects against anoikis by increasing BCL-xL expression levels. J Biol Chem 281: 17758–17767. - PubMed
-
- Asangani IA, Rasheed SA, Nikolova DA, Leupold JH, Colburn NH, Post S, Allgayer H. 2008. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 27: 2128–2136. - PubMed
-
- Baraniskin A, Birkenkamp-Demtroder K, Maghnouj A, Zollner H, Munding J, Klein-Scory S, Reinacher-Schick A, Schwarte-Waldhoff I, Schmiegel W, Hahn SA. 2012. MiR-30a-5p suppresses tumor growth in colon carcinoma by targeting DTL. Carcinogenesis 33: 732–739. - PubMed
-
- Berezikov E. 2011. Evolution of microRNA diversity and regulation in animals. Nat Rev 12: 846–860. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
