Optimization of routine KRAS mutation PCR-based testing procedure for rational individualized first-line-targeted therapy selection in metastatic colorectal cancer
- PMID: 24133623
- PMCID: PMC3797557
- DOI: 10.1002/cam4.47
Optimization of routine KRAS mutation PCR-based testing procedure for rational individualized first-line-targeted therapy selection in metastatic colorectal cancer
Abstract
KRAS mutation detection represents a crucial issue in metastatic colorectal cancer (mCRC). The optimization of KRAS mutation detection delay enabling rational prescription of first-line treatment in mCRC including anti-EGFR-targeted therapy requires robust and rapid molecular biology techniques. Routine analysis of mutations in codons 12 and 13 on 674 paraffin-embedded tissue specimens of mCRC has been performed for KRAS mutations detection using three molecular biology techniques, that is, high-resolution melting (HRM), polymerase chain reaction restriction fragment length polymorphism (PCR-RFLP), and allelic discrimination PCR (TaqMan PCR). Discordant cases were assessed with COBAS 4800 KRAS CE-IVD assay. Among the 674 tumor specimens, 1.5% (10/674) had excessive DNA degradation and could not be analyzed. KRAS mutations were detected in 38.0% (256/674) of the analysable specimens (82.4% in codon 12 and 17.6% in codon 13). Among 613 specimens in whom all three techniques were used, 12 (2.0%) cases of discordance between the three techniques were observed. 83.3% (10/12) of the discordances were due to PCR-RFLP as confirmed by COBAS 4800 retrospective analysis. The three techniques were statistically comparable (κ > 0.9; P < 0.001). From these results, optimization of the routine procedure consisted of proceeding to systematic KRAS detection using HRM and TaqMan and PCR-RFLP in case of discordance and allowed significant decrease in delays. The results showed an excellent correlation between the three techniques. Using HRM and TaqMan warrants high-quality and rapid-routine KRAS mutation detection in paraffin-embedded tumor specimens. The new procedure allowed a significant decrease in delays for reporting results, enabling rational prescription of first-line-targeted therapy in mCRC.
Keywords: Colorectal cancer; HRM; KRAS; PCR-RFLP; TaqMan PCR.
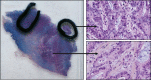
Figures


Similar articles
-
Comparison of COBAS 4800 KRAS, TaqMan PCR and high resolution melting PCR assays for the detection of KRAS somatic mutations in formalin-fixed paraffin embedded colorectal carcinomas.Virchows Arch. 2013 Mar;462(3):329-35. doi: 10.1007/s00428-013-1380-x. Epub 2013 Feb 12. Virchows Arch. 2013. PMID: 23400679
-
Sensitive multiplex detection of KRAS codons 12 and 13 mutations in paraffin-embedded tissue specimens.J Clin Pathol. 2011 Jan;64(1):30-6. doi: 10.1136/jcp.2010.081539. Epub 2010 Oct 28. J Clin Pathol. 2011. PMID: 21030527
-
KRAS and BRAF mutation analysis in routine molecular diagnostics: comparison of three testing methods on formalin-fixed, paraffin-embedded tumor-derived DNA.J Mol Diagn. 2012 May-Jun;14(3):247-55. doi: 10.1016/j.jmoldx.2012.01.011. Epub 2012 Mar 14. J Mol Diagn. 2012. PMID: 22425762
-
KRAS mutation testing in human cancers: The pathologist's role in the era of personalized medicine.Adv Anat Pathol. 2010 Jan;17(1):23-32. doi: 10.1097/PAP.0b013e3181c6962f. Adv Anat Pathol. 2010. PMID: 20032635 Review.
-
KRAS mutation testing in colorectal cancer as an example of the pathologist's role in personalized targeted therapy: a practical approach.Pol J Pathol. 2012 Nov;63(3):145-64. doi: 10.5114/pjp.2012.31499. Pol J Pathol. 2012. PMID: 23161231 Review.
Cited by
-
Evaluation and validation of HPV real-time PCR assay for the detection of HPV DNA in oral cytobrush and FFPE samples.Sci Rep. 2018 Jul 27;8(1):11313. doi: 10.1038/s41598-018-29790-z. Sci Rep. 2018. PMID: 30054550 Free PMC article.
-
Molecular Genetic Techniques in Biomarker Analysis Relevant for Drugs Centrally Approved in Europe.Mol Diagn Ther. 2022 Jan;26(1):89-103. doi: 10.1007/s40291-021-00567-x. Epub 2021 Dec 14. Mol Diagn Ther. 2022. PMID: 34905151 Free PMC article. Review.
-
Overview of Molecular Diagnostics in Irish Clinical Oncology.HRB Open Res. 2025 Jun 9;7:16. doi: 10.12688/hrbopenres.13822.2. eCollection 2024. HRB Open Res. 2025. PMID: 40687499 Free PMC article.
-
Association between proto-oncogene mutations and clinicopathologic characteristics and overall survival in colorectal cancer in East Azerbaijan, Iran.Onco Targets Ther. 2016 Dec 7;9:7385-7395. doi: 10.2147/OTT.S116373. eCollection 2016. Onco Targets Ther. 2016. PMID: 27994469 Free PMC article.
-
Spatial heterogeneity of KRAS mutations in colorectal cancers in northern France.Cancer Manag Res. 2019 Sep 13;11:8337-8344. doi: 10.2147/CMAR.S211119. eCollection 2019. Cancer Manag Res. 2019. PMID: 31571990 Free PMC article.
References
-
- Jemal A., Bray F., Center M. M., Ferlay J., Ward E., Forman D. Global cancer statistics. CA Cancer J. Clin. 2011;61:69–90. - PubMed
-
- Amado R. G., Wolf M., Peeters M., Van Cutsem E., Siena S., Freeman D. J. Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J. Clin. Oncol. 2008;26:1626–1634. - PubMed
-
- De Roock W., Piessevaux H., De Schutter J., Janssens M., De Hertogh G., Personeni N. KRAS wild-type state predicts survival and is associated to early radiological response in metastatic colorectal cancer treated with cetuximab. Ann. Oncol. 2008;19:508–515. - PubMed
-
- Lievre A., Bachet J. B., Boige V., Cayre A., Le Corre D., Buc E. KRAS mutations as an independent prognostic factor in patients with advanced colorectal cancer treated with cetuximab. J. Clin. Oncol. 2008;26:374–379. - PubMed
-
- Lievre A., Bachet J. B., Le Corre D., Boige V., Landi B., Emile J. F. KRAS mutation status is predictive of response to cetuximab therapy in colorectal cancer. Cancer Res. 2006;66:3992–3995. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

