Genome-level analyses of Mycobacterium bovis lineages reveal the role of SNPs and antisense transcription in differential gene expression
- PMID: 24134787
- PMCID: PMC3856593
- DOI: 10.1186/1471-2164-14-710
Genome-level analyses of Mycobacterium bovis lineages reveal the role of SNPs and antisense transcription in differential gene expression
Abstract
Background: Bovine tuberculosis (bTB) is a disease with major implications for animal welfare and productivity, as well as having the potential for zoonotic transmission. In Great Britain (GB) alone, controlling bTB costs in the region of £ 100 million annually, with the current control scheme seemingly unable to stop the inexorable spread of infection. One aspect that may be driving the epidemic is evolution of the causative pathogen, Mycobacterium bovis. To understand the underlying genetic changes that may be responsible for this evolution, we performed a comprehensive genome-level analyses of 4 M. bovis strains that encompass the main molecular types of the pathogen circulating in GB.
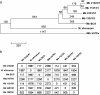
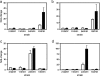
Results: We have used a combination of genome sequencing, transcriptome analyses, and recombinant DNA technology to define genetic differences across the major M. bovis lineages circulating in GB that may give rise to phenotypic differences of practical importance. The genomes of three M. bovis field isolates were sequenced using Illumina sequencing technology and strain specific differences in gene expression were measured during in vitro growth and in ex vivo bovine alveolar macrophages using a whole genome amplicon microarray and a whole genome tiled oligonucleotide microarray. SNP/small base pair insertion and deletions and gene expression data were overlaid onto the genomic sequence of the fully sequenced strain of M. bovis 2122/97 to link observed strain specific genomic differences with differences in RNA expression.
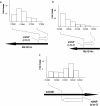
Conclusions: We show that while these strains show extensive similarities in their genetic make-up and gene expression profiles, they exhibit distinct expression of a subset of genes. We provide genomic, transcriptomic and functional data to show that synonymous point mutations (sSNPs) on the coding strand can lead to the expression of antisense transcripts on the opposing strand, a finding with implications for how we define a 'silent' nucleotide change. Furthermore, we show that transcriptomic data based solely on amplicon arrays can generate spurious results in terms of gene expression profiles due to hybridisation of antisense transcripts. Overall our data suggest that subtle genetic differences, such as sSNPS, may have important consequences for gene expression and subsequent phenotype.
Figures

Similar articles
-
Whole genome sequencing of the monomorphic pathogen Mycobacterium bovis reveals local differentiation of cattle clinical isolates.BMC Genomics. 2018 Jan 2;19(1):2. doi: 10.1186/s12864-017-4249-6. BMC Genomics. 2018. PMID: 29291727 Free PMC article.
-
Whole-transcriptome, high-throughput RNA sequence analysis of the bovine macrophage response to Mycobacterium bovis infection in vitro.BMC Genomics. 2013 Apr 8;14:230. doi: 10.1186/1471-2164-14-230. BMC Genomics. 2013. PMID: 23565803 Free PMC article.
-
Genetic profiling of Mycobacterium bovis strains from slaughtered cattle in Eritrea.PLoS Negl Trop Dis. 2018 Apr 17;12(4):e0006406. doi: 10.1371/journal.pntd.0006406. eCollection 2018 Apr. PLoS Negl Trop Dis. 2018. PMID: 29664901 Free PMC article.
-
Molecular Epidemiology of Mycobacterium bovis in Humans and Cattle.Zoonoses Public Health. 2016 Jun;63(4):251-64. doi: 10.1111/zph.12242. Epub 2015 Dec 18. Zoonoses Public Health. 2016. PMID: 26684712 Review.
-
Recent advances in our knowledge of Mycobacterium bovis: a feeling for the organism.Vet Microbiol. 2006 Feb 25;112(2-4):127-39. doi: 10.1016/j.vetmic.2005.11.050. Epub 2005 Dec 27. Vet Microbiol. 2006. PMID: 16384663 Review.
Cited by
-
The open pan-genome architecture and virulence landscape of Mycobacterium bovis.Microb Genom. 2021 Oct;7(10):000664. doi: 10.1099/mgen.0.000664. Microb Genom. 2021. PMID: 34714230 Free PMC article.
-
Deciphering the metabolic response of Mycobacterium tuberculosis to nitrogen stress.Mol Microbiol. 2015 Sep;97(6):1142-57. doi: 10.1111/mmi.13091. Epub 2015 Jul 17. Mol Microbiol. 2015. PMID: 26077160 Free PMC article.
-
Primary Role of Suppressor of Cytokine Signaling 1 in Mycobacterium bovis BCG Infection.Infect Immun. 2018 Oct 25;86(11):e00376-18. doi: 10.1128/IAI.00376-18. Print 2018 Nov. Infect Immun. 2018. PMID: 30181351 Free PMC article.
-
Small RNAs in mycobacteria: an unfolding story.Front Cell Infect Microbiol. 2014 Jul 24;4:96. doi: 10.3389/fcimb.2014.00096. eCollection 2014. Front Cell Infect Microbiol. 2014. PMID: 25105095 Free PMC article. Review.
-
Genome-wide SNP and InDel mutations in Mycobacterium tuberculosis associated with rifampicin and isoniazid resistance.Int J Clin Exp Pathol. 2018 Aug 1;11(8):3903-3914. eCollection 2018. Int J Clin Exp Pathol. 2018. PMID: 31949778 Free PMC article.
References
-
- Monaghan ML, Doherty ML, Collins JD, Kazda JF, Quinn PJ. The tuberculin test. Vet Microbiol. 1994;40(1–2):111–124. - PubMed
-
- Frothingham R, Meeker-O’Connell WA. Genetic diversity in the Mycobacterium tuberculosis complex based on variable numbers of tandem DNA repeats. Microbiology. 1998;144(Pt 5):1189–1196. - PubMed
-
- Winder CL, Gordon SV, Dale J, Hewinson RG, Goodacre R. Metabolic fingerprints of Mycobacterium bovis cluster with molecular type: implications for genotype-phenotype links. Microbiology. 2006;152(Pt 9):2757–2765. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

