Extensive variation in chromatin states across humans
- PMID: 24136358
- PMCID: PMC4075767
- DOI: 10.1126/science.1242510
Extensive variation in chromatin states across humans
Abstract
The majority of disease-associated variants lie outside protein-coding regions, suggesting a link between variation in regulatory regions and disease predisposition. We studied differences in chromatin states using five histone modifications, cohesin, and CTCF in lymphoblastoid lines from 19 individuals of diverse ancestry. We found extensive signal variation in regulatory regions, which often switch between active and repressed states across individuals. Enhancer activity is particularly diverse among individuals, whereas gene expression remains relatively stable. Chromatin variability shows genetic inheritance in trios, correlates with genetic variation and population divergence, and is associated with disruptions of transcription factor binding motifs. Overall, our results provide insights into chromatin variation among humans.
Figures


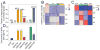

Comment in
-
Gene regulation: from genetic variation to phenotype via chromatin.Nat Rev Genet. 2013 Dec;14(12):824. doi: 10.1038/nrg3622. Epub 2013 Oct 29. Nat Rev Genet. 2013. PMID: 24166029 No abstract available.
-
Genetics. Genetics driving epigenetics.Science. 2013 Nov 8;342(6159):705-6. doi: 10.1126/science.1246755. Science. 2013. PMID: 24202168 No abstract available.
References
-
- Wellcome Trust Case Control Consortium. Nature. 2007;447:661–678.
-
- McCarthy MI, et al. Nat Rev Genet. 2008;9:356–369. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

