Quantitative imaging of transcription in living Drosophila embryos links polymerase activity to patterning
- PMID: 24139738
- PMCID: PMC3828032
- DOI: 10.1016/j.cub.2013.08.054
Quantitative imaging of transcription in living Drosophila embryos links polymerase activity to patterning
Abstract
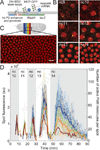
Spatiotemporal patterns of gene expression are fundamental to every developmental program. The resulting macroscopic domains have been mainly characterized by their levels of gene products. However, the establishment of such patterns results from differences in the dynamics of microscopic events in individual cells such as transcription. It is unclear how these microscopic decisions lead to macroscopic patterns, as measurements in fixed tissue cannot access the underlying transcriptional dynamics. In vivo transcriptional dynamics have long been approached in single-celled organisms, but never in a multicellular developmental context. Here, we directly address how boundaries of gene expression emerge in the Drosophila embryo by measuring the absolute number of actively transcribing polymerases in real time in individual nuclei. Specifically, we show that the formation of a boundary cannot be quantitatively explained by the rate of mRNA production in each cell, but instead requires amplification of the dynamic range of the expression boundary. This amplification is accomplished by nuclei randomly adopting active or inactive states of transcription, leading to a collective effect where the fraction of active nuclei is modulated in space. Thus, developmental patterns are not just the consequence of reproducible transcriptional dynamics in individual nuclei, but are the result of averaging expression over space and time.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures



Comment in
-
Development: seeing the pattern.Nat Rev Genet. 2013 Dec;14(12):823. doi: 10.1038/nrg3619. Epub 2013 Oct 29. Nat Rev Genet. 2013. PMID: 24166030 No abstract available.
-
Development: lights, camera, action--the Drosophila embryo goes live!Curr Biol. 2013 Nov 4;23(21):R965-7. doi: 10.1016/j.cub.2013.09.054. Curr Biol. 2013. PMID: 24200326
References
-
- Poustelnikova E, Pisarev A, Blagov M, Samsonova M, Reinitz J. A database for management of gene expression data in situ. Bioinformatics. 2004;20:2212–2221. - PubMed
-
- Jaeger J, Surkova S, Blagov M, Janssens H, Kosman D, Kozlov KN, Manu, Myasnikova E, Vanario-Alonso CE, Samsonova M, et al. Dynamic control of positional information in the early Drosophila embryo. Nature. 2004;430:368–371. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

