Mechanism of allosteric activation of SAMHD1 by dGTP
- PMID: 24141705
- PMCID: PMC3833828
- DOI: 10.1038/nsmb.2692
Mechanism of allosteric activation of SAMHD1 by dGTP
Abstract
SAMHD1, a dNTP triphosphohydrolase (dNTPase), has a key role in human innate immunity. It inhibits infection of blood cells by retroviruses, including HIV, and prevents the development of the autoinflammatory Aicardi-Goutières syndrome (AGS). The inactive apo-SAMHD1 interconverts between monomers and dimers, and in the presence of dGTP the protein assembles into catalytically active tetramers. Here, we present the crystal structure of the human tetrameric SAMHD1-dGTP complex. The structure reveals an elegant allosteric mechanism of activation through dGTP-induced tetramerization of two inactive dimers. Binding of dGTP to four allosteric sites promotes tetramerization and induces a conformational change in the substrate-binding pocket to yield the catalytically active enzyme. Structure-based biochemical and cell-based biological assays confirmed the proposed mechanism. The SAMHD1 tetramer structure provides the basis for a mechanistic understanding of its function in HIV restriction and the pathogenesis of AGS.
Figures


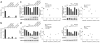

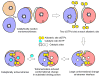
References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

