ubiJ, a new gene required for aerobic growth and proliferation in macrophage, is involved in coenzyme Q biosynthesis in Escherichia coli and Salmonella enterica serovar Typhimurium
- PMID: 24142253
- PMCID: PMC3911131
- DOI: 10.1128/JB.01065-13
ubiJ, a new gene required for aerobic growth and proliferation in macrophage, is involved in coenzyme Q biosynthesis in Escherichia coli and Salmonella enterica serovar Typhimurium
Abstract
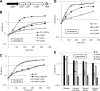
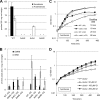
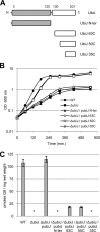
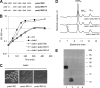
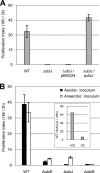
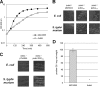
Ubiquinone (coenzyme Q or Q8) is a redox active lipid which functions in the respiratory electron transport chain and plays a crucial role in energy-generating processes. In both Escherichia coli and Salmonella enterica serovar Typhimurium, the yigP gene is located between ubiE and ubiB, all three being likely to constitute an operon. In this work, we showed that the uncharacterized yigP gene was involved in Q8 biosynthesis in both strains, and we have renamed it ubiJ. Under aerobic conditions, an ubiJ mutant was found to be impaired for Q8 biosynthesis and for growth in rich medium but did not present any defect anaerobically. Surprisingly, the C-terminal 50 amino acids, predicted to interact with lipids, were sufficient to restore Q8 biosynthesis and growth of the ubiJ mutant. Salmonella ubiE and ubiB mutants were impaired in Q8 biosynthesis and in respiration using different electron acceptors. Moreover, ubiE, ubiJ, and ubiB mutants were all impaired for Salmonella intracellular proliferation in macrophages. Taken together, our data establish an important role for UbiJ in Q8 biosynthesis and reveal an unexpected link between Q8 and virulence. They also emphasize that Salmonella organisms in an intracellular lifestyle rely on aerobic respiration to survive and proliferate within macrophages.
Figures
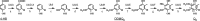
References
-
- Wallace BJ, Young IG. 1977. Role of quinones in electron transport to oxygen and nitrate in Escherichia coli. Studies with a ubiA− menA− double quinone mutant. Biochim. Biophys. Acta 461:84–100 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

