LIN28B promotes growth and tumorigenesis of the intestinal epithelium via Let-7
- PMID: 24142874
- PMCID: PMC3814644
- DOI: 10.1101/gad.224659.113
LIN28B promotes growth and tumorigenesis of the intestinal epithelium via Let-7
Abstract
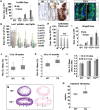
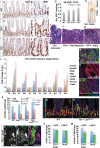
The RNA-binding proteins LIN28A and LIN28B have diverse functions in embryonic stem cells, cellular reprogramming, growth, and oncogenesis. Many of these effects occur via direct inhibition of Let-7 microRNAs (miRNAs), although Let-7-independent effects have been surmised. We report that intestine targeted expression of LIN28B causes intestinal hypertrophy, crypt expansion, and Paneth cell loss. Furthermore, LIN28B fosters intestinal polyp and adenocarcinoma formation. To examine potential Let-7-independent functions of LIN28B, we pursued ribonucleoprotein cross-linking, immunoprecipitation, and high-throughput sequencing (CLIP-seq) to identify direct RNA targets. This revealed that LIN28B bound a substantial number of mRNAs and modestly augmented protein levels of these target mRNAs in vivo. Conversely, Let-7 had a profound effect; modulation of Let-7 levels via deletion of the mirLet7c2/mirLet7b genes recapitulated effects of Lin28b overexpression. Furthermore, intestine-specific Let-7 expression could reverse hypertrophy and Paneth cell depletion caused by Lin28b. This was independent of effects on insulin-PI3K-mTOR signaling. Our study reveals that Let-7 miRNAs are critical for repressing intestinal tissue growth and promoting Paneth cell differentiation. Let-7-dependent effects of LIN28B may supersede Let-7-independent effects on intestinal tissue growth. In summary, LIN28B can definitively act as an oncogene in the absence of canonical genetic alterations.
Keywords: CLIP-seq; LIN28B; Let-7; Paneth cells; colon cancer; intestinal epithelium.
Figures





References
-
- Barker N, Clevers H 2010. Leucine-rich repeat-containing G-protein-coupled receptors as markers of adult stem cells. Gastroenterology 138: 1681–1696 - PubMed
-
- Brooks DG, James RM, Patek CE, Williamson J, Arends MJ 2001. Mutant K-ras enhances apoptosis in embryonic stem cells in combination with DNA damage and is associated with increased levels of p19(ARF). Oncogene 20: 2144–2152 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P50 CA095103/CA/NCI NIH HHS/United States
- P30 DK050306/DK/NIDDK NIH HHS/United States
- DK083111/DK/NIDDK NIH HHS/United States
- R01 DK056645/DK/NIDDK NIH HHS/United States
- K01DK093885/DK/NIDDK NIH HHS/United States
- K01 DK093885/DK/NIDDK NIH HHS/United States
- DP2 DK083111/DK/NIDDK NIH HHS/United States
- P30 CA068485/CA/NCI NIH HHS/United States
- P30 CA016520/CA/NCI NIH HHS/United States
- DK083355/DK/NIDDK NIH HHS/United States
- P30-DK050306/DK/NIDDK NIH HHS/United States
- R01 DK083355/DK/NIDDK NIH HHS/United States
- R01-DK056645/DK/NIDDK NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
