Identification of genes potentially regulated by human polynucleotide phosphorylase (hPNPase old-35) using melanoma as a model
- PMID: 24143183
- PMCID: PMC3797080
- DOI: 10.1371/journal.pone.0076284
Identification of genes potentially regulated by human polynucleotide phosphorylase (hPNPase old-35) using melanoma as a model
Abstract
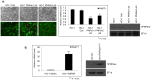
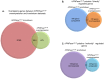
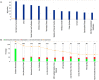
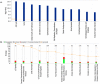
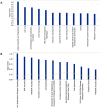
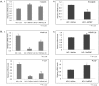
Human Polynucleotide Phosphorylase (hPNPase(old-35) or PNPT1) is an evolutionarily conserved 3'→ 5' exoribonuclease implicated in the regulation of numerous physiological processes including maintenance of mitochondrial homeostasis, mtRNA import and aging-associated inflammation. From an RNase perspective, little is known about the RNA or miRNA species it targets for degradation or whose expression it regulates; except for c-myc and miR-221. To further elucidate the functional implications of hPNPase(old-35) in cellular physiology, we knocked-down and overexpressed hPNPase(old-35) in human melanoma cells and performed gene expression analyses to identify differentially expressed transcripts. Ingenuity Pathway Analysis indicated that knockdown of hPNPase(old-35) resulted in significant gene expression changes associated with mitochondrial dysfunction and cholesterol biosynthesis; whereas overexpression of hPNPase(old-35) caused global changes in cell-cycle related functions. Additionally, comparative gene expression analyses between our hPNPase(old-35) knockdown and overexpression datasets allowed us to identify 77 potential "direct" and 61 potential "indirect" targets of hPNPase(old-35) which formed correlated networks enriched for cell-cycle and wound healing functional association, respectively. These results provide a comprehensive database of genes responsive to hPNPase(old-35) expression levels; along with the identification new potential candidate genes offering fresh insight into cellular pathways regulated by PNPT1 and which may be used in the future for possible therapeutic intervention in mitochondrial- or inflammation-associated disease phenotypes.
Conflict of interest statement
Figures



Similar articles
-
Analysis of global changes in gene expression induced by human polynucleotide phosphorylase (hPNPase(old-35)).J Cell Physiol. 2014 Dec;229(12):1952-62. doi: 10.1002/jcp.24645. J Cell Physiol. 2014. PMID: 24729470 Free PMC article.
-
Expression regulation and genomic organization of human polynucleotide phosphorylase, hPNPase(old-35), a Type I interferon inducible early response gene.Gene. 2003 Oct 16;316:143-56. doi: 10.1016/s0378-1119(03)00752-2. Gene. 2003. PMID: 14563561
-
Human polynucleotide phosphorylase selectively and preferentially degrades microRNA-221 in human melanoma cells.Proc Natl Acad Sci U S A. 2010 Jun 29;107(26):11948-53. doi: 10.1073/pnas.0914143107. Epub 2010 Jun 14. Proc Natl Acad Sci U S A. 2010. PMID: 20547861 Free PMC article.
-
Activity and Function in Human Cells of the Evolutionary Conserved Exonuclease Polynucleotide Phosphorylase.Int J Mol Sci. 2022 Jan 31;23(3):1652. doi: 10.3390/ijms23031652. Int J Mol Sci. 2022. PMID: 35163574 Free PMC article. Review.
-
Human polynucleotide phosphorylase (hPNPase old-35): an RNA degradation enzyme with pleiotrophic biological effects.Cell Cycle. 2006 May;5(10):1080-4. doi: 10.4161/cc.5.10.2741. Epub 2006 May 15. Cell Cycle. 2006. PMID: 16687933 Review.
Cited by
-
Identification of RNA processing factor LSM5 as a new adverse biomarker in nasopharyngeal carcinoma.Sci Rep. 2025 Mar 22;15(1):9901. doi: 10.1038/s41598-025-94968-1. Sci Rep. 2025. PMID: 40121338 Free PMC article.
-
Whole-exome sequencing identifies novel variants in PNPT1 causing oxidative phosphorylation defects and severe multisystem disease.Eur J Hum Genet. 2017 Jan;25(1):79-84. doi: 10.1038/ejhg.2016.128. Epub 2016 Oct 19. Eur J Hum Genet. 2017. PMID: 27759031 Free PMC article.
-
PNPase knockout results in mtDNA loss and an altered metabolic gene expression program.PLoS One. 2018 Jul 19;13(7):e0200925. doi: 10.1371/journal.pone.0200925. eCollection 2018. PLoS One. 2018. PMID: 30024931 Free PMC article.
-
Analysis of global changes in gene expression induced by human polynucleotide phosphorylase (hPNPase(old-35)).J Cell Physiol. 2014 Dec;229(12):1952-62. doi: 10.1002/jcp.24645. J Cell Physiol. 2014. PMID: 24729470 Free PMC article.
-
Examination of Epigenetic and other Molecular Factors Associated with mda-9/Syntenin Dysregulation in Cancer Through Integrated Analyses of Public Genomic Datasets.Adv Cancer Res. 2015;127:49-121. doi: 10.1016/bs.acr.2015.04.006. Epub 2015 May 23. Adv Cancer Res. 2015. PMID: 26093898 Free PMC article. Review.
References
-
- Arraiano C, Andrade J, Domingues S, Guinote I, Malecki M, et al. (2010) The critical role of RNA processing and degradation in the control of gene expression. FEMS Microbiol Rev 34: 883–923. - PubMed
-
- Deutscher MP, Li Z (2001) Exoribonucleases and their multiple roles in RNA metabolism. Prog Nucleic Acid Res Mol Biol 66: 67–105. - PubMed
-
- Deutscher MP (1993) Ribonuclease multiplicity, diversity, and complexity. J Biol Chem 268: 13011–13014. - PubMed
-
- Andrade J, Pobre V, Silva I, Domingues S, Arraiano C (2009) The role of 3′-5′ exoribonucleases in RNA degradation. Prog Mol Biol Transl Sci 85: 187–229. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

