Lanthanide-based imaging of protein-protein interactions in live cells
- PMID: 24144069
- PMCID: PMC3944735
- DOI: 10.1021/ic4018739
Lanthanide-based imaging of protein-protein interactions in live cells
Abstract
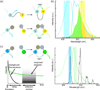
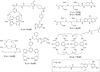
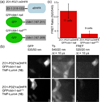
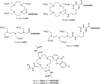
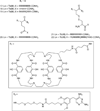
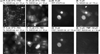
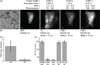
In order to deduce the molecular mechanisms of biological function, it is necessary to monitor changes in the subcellular location, activation, and interaction of proteins within living cells in real time. Förster resonance energy-transfer (FRET)-based biosensors that incorporate genetically encoded, fluorescent proteins permit high spatial resolution imaging of protein-protein interactions or protein conformational dynamics. However, a nonspecific fluorescence background often obscures small FRET signal changes, and intensity-based biosensor measurements require careful interpretation and several control experiments. These problems can be overcome by using lanthanide [Tb(III) or Eu(III)] complexes as donors and green fluorescent protein (GFP) or other conventional fluorophores as acceptors. Essential features of this approach are the long-lifetime (approximately milliseconds) luminescence of Tb(III) complexes and time-gated luminescence microscopy. This allows pulsed excitation, followed by a brief delay, which eliminates nonspecific fluorescence before the detection of Tb(III)-to-GFP emission. The challenges of intracellular delivery, selective protein labeling, and time-gated imaging of lanthanide luminescence are presented, and recent efforts to investigate the cellular uptake of lanthanide probes are reviewed. Data are presented showing that conjugation to arginine-rich, cell-penetrating peptides (CPPs) can be used as a general strategy for the cellular delivery of membrane-impermeable lanthanide complexes. A heterodimer of a luminescent Tb(III) complex, Lumi4, linked to trimethoprim and conjugated to nonaarginine via a reducible disulfide linker rapidly (∼10 min) translocates into the cytoplasm of Maden Darby canine kidney cells from the culture medium. With this reagent, the intracellular interaction between GFP fused to FK506 binding protein 12 (GFP-FKBP12) and the rapamycin binding domain of mTOR fused to Escherichia coli dihydrofolate reductase (FRB-eDHFR) were imaged at high signal-to-noise ratio with fast (1-3 s) image acquisition using a time-gated luminescence microscope. The data reviewed and presented here show that lanthanide biosensors enable fast, sensitive, and technically simple imaging of protein-protein interactions in live cells.
Figures

Similar articles
-
Lanthanide-based resonance energy transfer biosensors for live-cell applications.Methods Enzymol. 2021;651:291-311. doi: 10.1016/bs.mie.2021.01.010. Epub 2021 Feb 23. Methods Enzymol. 2021. PMID: 33888207
-
Evaluating the performance of time-gated live-cell microscopy with lanthanide probes.Biophys J. 2015 Jul 21;109(2):240-8. doi: 10.1016/j.bpj.2015.06.028. Biophys J. 2015. PMID: 26200860 Free PMC article.
-
Cytoplasmic delivery and selective, multicomponent labeling with oligoarginine-linked protein tags.Bioconjug Chem. 2015 Mar 18;26(3):460-5. doi: 10.1021/bc500550z. Epub 2015 Feb 24. Bioconjug Chem. 2015. PMID: 25675354 Free PMC article.
-
Development of responsive lanthanide-based magnetic resonance imaging and luminescent probes for biological applications.Chem Pharm Bull (Tokyo). 2010 Oct;58(10):1283-94. doi: 10.1248/cpb.58.1283. Chem Pharm Bull (Tokyo). 2010. PMID: 20930392 Review.
-
Luminescent lanthanide metallopeptides for biomolecule sensing and cellular imaging.Chem Commun (Camb). 2024 Oct 29;60(87):12650-12661. doi: 10.1039/d4cc03205e. Chem Commun (Camb). 2024. PMID: 39327864 Free PMC article. Review.
Cited by
-
Luminescence resonance energy transfer between genetically encoded donor and acceptor for protein-protein interaction studies in the molecular chaperone HSP70/HSP90 complexes.Sci Rep. 2018 Feb 12;8(1):2801. doi: 10.1038/s41598-018-21210-6. Sci Rep. 2018. PMID: 29434293 Free PMC article.
-
Time-gated FRET nanoassemblies for rapid and sensitive intra- and extracellular fluorescence imaging.Sci Adv. 2016 Jun 10;2(6):e1600265. doi: 10.1126/sciadv.1600265. eCollection 2016 Jun. Sci Adv. 2016. PMID: 27386579 Free PMC article.
-
Smart Applications of Lanthanide Chelates-based Luminescent Probes in Bio-imaging.Mini Rev Med Chem. 2025;25(7):505-520. doi: 10.2174/0113895575350677250101060606. Mini Rev Med Chem. 2025. PMID: 39886779 Review.
-
Biosensing Tacrolimus in Human Whole Blood by Using a Drug Receptor Fused to the Emerald Green Fluorescent Protein.Anal Chem. 2022 Nov 29;94(47):16337-16344. doi: 10.1021/acs.analchem.2c03122. Epub 2022 Nov 16. Anal Chem. 2022. PMID: 36382944 Free PMC article.
-
Recent Advances in Luminescence Imaging of Biological Systems Using Lanthanide(III) Luminescent Complexes.Molecules. 2020 Apr 29;25(9):2089. doi: 10.3390/molecules25092089. Molecules. 2020. PMID: 32365719 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

