Effects of prostaglandin E2 on p53 mRNA transcription and p53 mutagenesis during T-cell-independent human B-cell clonal expansion
- PMID: 24145719
- PMCID: PMC3898655
- DOI: 10.1096/fj.13-237792
Effects of prostaglandin E2 on p53 mRNA transcription and p53 mutagenesis during T-cell-independent human B-cell clonal expansion
Abstract
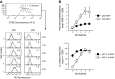
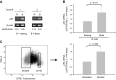
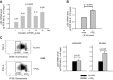
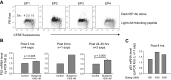
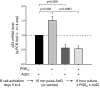
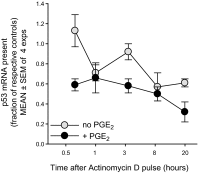
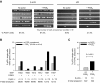
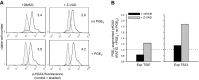
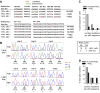
Within T-cell-dependent germinal centers, p53 gene transcription is repressed by Bcl-6 and is thus less vulnerable to mutation. Malignant lymphomas within inflamed extranodal sites exhibit a relatively high incidence of p53 mutations. The latter might originate from normal B-cell clones manifesting activation-induced cytosine deaminase (AID) and up-regulated p53 following T-cell-independent (TI) stimulation. We here examine p53 gene transcription in such TI clones, with a focus on modulatory effects of prostaglandin E2 (PGE2), and evaluate progeny for p53 mutations. Resting IgM(+)IgD(+)CD27(-) B cells from human tonsils were labeled with CFSE and stimulated in vitro with complement-coated antigen surrogate, IL-4, and BAFF ± exogenous PGE2 (50 nM) or an analog specific for the EP2 PGE2 receptor. We use flow cytometry to measure p53 and AID protein within variably divided blasts, qRT-PCR of p53 mRNA from cultures with or without actinomycin D to monitor mRNA transcription/stability, and single-cell p53 RT-PCR/sequencing to assess progeny for p53 mutations. We report that EP2 signaling triggers increased p53 gene transcriptional activity in AID(+) cycling blasts (P<0.01). Progeny exhibit p53 mutations at a frequency (8.5 × 10(-4)) greater than the baseline error rate (<0.8 × 10(-4)). We conclude that, devoid of the repressive influences of Bcl-6, dividing B lymphoblasts in inflamed tissues should display heightened p53 transcription and increased risk of p53 mutagenesis.
Keywords: B lymphocyte; TP53; eicosanoid; inflammation; lymphoma.
Figures
Similar articles
-
A p53 axis regulates B cell receptor-triggered, innate immune system-driven B cell clonal expansion.J Immunol. 2012 Jun 15;188(12):6093-108. doi: 10.4049/jimmunol.1103037. Epub 2012 May 18. J Immunol. 2012. PMID: 22611237 Free PMC article.
-
A cyclooxygenase-2/prostaglandin E2 pathway augments activation-induced cytosine deaminase expression within replicating human B cells.J Immunol. 2010 Nov 1;185(9):5300-14. doi: 10.4049/jimmunol.1000574. Epub 2010 Oct 4. J Immunol. 2010. PMID: 20921530
-
EP4/EP2 receptor-specific prostaglandin E2 regulation of interleukin-6 generation by human HSB.2 early T cells.J Pharmacol Exp Ther. 1998 Sep;286(3):1420-6. J Pharmacol Exp Ther. 1998. PMID: 9732406
-
Prostaglandin E2 differentially modulates IL-5 gene expression in activated human T lymphocytes depending on the costimulatory signal.J Allergy Clin Immunol. 1998 Feb;101(2 Pt 1):231-40. doi: 10.1016/s0091-6749(98)70388-4. J Allergy Clin Immunol. 1998. PMID: 9500757
-
Prostaglandin E2 suppresses poly I: C-stimulated cytokine production via EP2 and EP3 in immortalized human corneal epithelial cells.Cornea. 2012 Nov;31(11):1294-8. doi: 10.1097/ICO.0b013e318242fd7c. Cornea. 2012. PMID: 22475642
Cited by
-
Mechanism for IL-15-Driven B Cell Chronic Lymphocytic Leukemia Cycling: Roles for AKT and STAT5 in Modulating Cyclin D2 and DNA Damage Response Proteins.J Immunol. 2019 May 15;202(10):2924-2944. doi: 10.4049/jimmunol.1801142. Epub 2019 Apr 15. J Immunol. 2019. PMID: 30988120 Free PMC article.
-
Sphingolipids and Lymphomas: A Double-Edged Sword.Cancers (Basel). 2022 Apr 19;14(9):2051. doi: 10.3390/cancers14092051. Cancers (Basel). 2022. PMID: 35565181 Free PMC article. Review.
-
TLR-9 and IL-15 Synergy Promotes the In Vitro Clonal Expansion of Chronic Lymphocytic Leukemia B Cells.J Immunol. 2015 Aug 1;195(3):901-23. doi: 10.4049/jimmunol.1403189. Epub 2015 Jul 1. J Immunol. 2015. PMID: 26136429 Free PMC article.
-
BCL2 Protein Progressively Declines during Robust CLL Clonal Expansion: Potential Impact on Venetoclax Clinical Efficacy and Insights on Mechanism.Lymphatics. 2024 Jun;2(2):50-78. doi: 10.3390/lymphatics2020005. Epub 2024 Mar 28. Lymphatics. 2024. PMID: 39664277 Free PMC article.
-
Exosomes molecular diagnostics: Direct conversion of exosomes into the cDNA for gene amplification by two-step polymerase chain reaction.J Biol Methods. 2018 Jul 25;5(3):e96. doi: 10.14440/jbm.2018.249. eCollection 2018. J Biol Methods. 2018. PMID: 31453246 Free PMC article.
References
-
- Lane D. P. (1992) Cancer p53, guardian of the genome. Nature 358, 15–16 - PubMed
-
- Phan R. T., Dalla-Favera R. (2004) The BCL6 proto-oncogene suppresses p53 expression in germinal-centre B cells. Nature 432, 635–639 - PubMed
-
- Bachl J., Carlson C., Gray-Schopfer V., Dessing M., Olsson C. (2001) Increased transcription levels induce higher mutation rates in a hypermutating cell line. J. Immunol. 166, 5051–5057 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

