Recurrent patterns of DNA methylation in the ZNF154, CASP8, and VHL promoters across a wide spectrum of human solid epithelial tumors and cancer cell lines
- PMID: 24149212
- PMCID: PMC3933495
- DOI: 10.4161/epi.26701
Recurrent patterns of DNA methylation in the ZNF154, CASP8, and VHL promoters across a wide spectrum of human solid epithelial tumors and cancer cell lines
Abstract
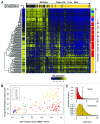
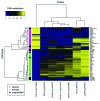
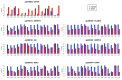
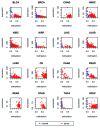
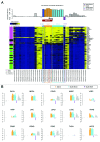
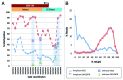
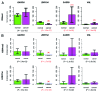
The study of aberrant DNA methylation in cancer holds the key to the discovery of novel biological markers for diagnostics and can help to delineate important mechanisms of disease. We have identified 12 loci that are differentially methylated in serous ovarian cancers and endometrioid ovarian and endometrial cancers with respect to normal control samples. The strongest signal showed hypermethylation in tumors at a CpG island within the ZNF154 promoter. We show that hypermethylation of this locus is recurrent across solid human epithelial tumor samples for 15 of 16 distinct cancer types from TCGA. Furthermore, ZNF154 hypermethylation is strikingly present across a diverse panel of ENCODE cell lines, but only in those derived from tumor cells. By extending our analysis from the Illumina 27K Infinium platform to the 450K platform, to sequencing of PCR amplicons from bisulfite treated DNA, we demonstrate that hypermethylation extends across the breadth of the ZNF154 CpG island. We have also identified recurrent hypomethylation in two genomic regions associated with CASP8 and VHL. These three genes exhibit significant negative correlation between methylation and gene expression across many cancer types, as well as patterns of DNaseI hypersensitivity and histone marks that reflect different chromatin accessibility in cancer vs. normal cell lines. Our findings emphasize hypermethylation of ZNF154 as a biological marker of relevance for tumor identification. Epigenetic modifications affecting the promoters of ZNF154, CASP8, and VHL are shared across a vast array of tumor types and may therefore be important for understanding the genomic landscape of cancer.
Keywords: CASP8; DNA methylation; VHL; ZNF154; cancer; chromatin; endometrioid endometrial cancer; endometrioid ovarian cancer; epigenetics; ovarian papillary serous tumors of low malignant potential; pan-cancer; serous ovarian cancer.
Figures

Similar articles
-
Robust Detection of DNA Hypermethylation of ZNF154 as a Pan-Cancer Locus with in Silico Modeling for Blood-Based Diagnostic Development.J Mol Diagn. 2016 Mar;18(2):283-98. doi: 10.1016/j.jmoldx.2015.11.004. Epub 2016 Feb 5. J Mol Diagn. 2016. PMID: 26857064 Free PMC article.
-
Epigenetic analysis of sporadic and Lynch-associated ovarian cancers reveals histology-specific patterns of DNA methylation.Epigenetics. 2014 Dec;9(12):1577-87. doi: 10.4161/15592294.2014.983374. Epigenetics. 2014. PMID: 25625843 Free PMC article.
-
Hypermethylation of miR-203 in endometrial carcinomas.Gynecol Oncol. 2014 May;133(2):340-5. doi: 10.1016/j.ygyno.2014.02.009. Epub 2014 Feb 14. Gynecol Oncol. 2014. PMID: 24530564 Free PMC article.
-
DNA methylation profiles in ovarian cancer: implication in diagnosis and therapy (Review).Mol Med Rep. 2014 Jul;10(1):3-9. doi: 10.3892/mmr.2014.2221. Epub 2014 May 8. Mol Med Rep. 2014. PMID: 24821107 Free PMC article. Review.
-
Epigenetics in ovarian cancer.Semin Cancer Biol. 2018 Aug;51:160-169. doi: 10.1016/j.semcancer.2017.08.003. Epub 2017 Aug 3. Semin Cancer Biol. 2018. PMID: 28782606 Free PMC article. Review.
Cited by
-
Robust Detection of DNA Hypermethylation of ZNF154 as a Pan-Cancer Locus with in Silico Modeling for Blood-Based Diagnostic Development.J Mol Diagn. 2016 Mar;18(2):283-98. doi: 10.1016/j.jmoldx.2015.11.004. Epub 2016 Feb 5. J Mol Diagn. 2016. PMID: 26857064 Free PMC article.
-
Long-range epigenetic regulation is conferred by genetic variation located at thousands of independent loci.Nat Commun. 2015 Feb 26;6:6326. doi: 10.1038/ncomms7326. Nat Commun. 2015. PMID: 25716334 Free PMC article.
-
Low Expression of ZNF154 is Related to Poor Prognosis in Gastric Cancer.Cancer Manag Res. 2022 Feb 17;14:659-672. doi: 10.2147/CMAR.S340053. eCollection 2022. Cancer Manag Res. 2022. PMID: 35210862 Free PMC article.
-
Genome-wide host methylation profiling of anal and cervical carcinoma.PLoS One. 2021 Dec 9;16(12):e0260857. doi: 10.1371/journal.pone.0260857. eCollection 2021. PLoS One. 2021. PMID: 34882728 Free PMC article.
-
DNA methylation marker to estimate ovarian cancer cell fraction.Med Oncol. 2022 Feb 23;39(5):78. doi: 10.1007/s12032-022-01679-y. Med Oncol. 2022. PMID: 35195779
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous
