Local label learning (LLL) for subcortical structure segmentation: application to hippocampus segmentation
- PMID: 24151008
- PMCID: PMC6869539
- DOI: 10.1002/hbm.22359
Local label learning (LLL) for subcortical structure segmentation: application to hippocampus segmentation
Abstract
Automatic and reliable segmentation of subcortical structures is an important but difficult task in quantitative brain image analysis. Multi-atlas based segmentation methods have attracted great interest due to their promising performance. Under the multi-atlas based segmentation framework, using deformation fields generated for registering atlas images onto a target image to be segmented, labels of the atlases are first propagated to the target image space and then fused to get the target image segmentation based on a label fusion strategy. While many label fusion strategies have been developed, most of these methods adopt predefined weighting models that are not necessarily optimal. In this study, we propose a novel local label learning strategy to estimate the target image's segmentation label using statistical machine learning techniques. In particular, we use a L1-regularized support vector machine (SVM) with a k nearest neighbor (kNN) based training sample selection strategy to learn a classifier for each of the target image voxel from its neighboring voxels in the atlases based on both image intensity and texture features. Our method has produced segmentation results consistently better than state-of-the-art label fusion methods in validation experiments on hippocampal segmentation of over 100 MR images obtained from publicly available and in-house datasets. Volumetric analysis has also demonstrated the capability of our method in detecting hippocampal volume changes due to Alzheimer's disease.
Keywords: SVM; hippocampal segmentation; local label learning; multi-atlas based segmentation.
Copyright © 2013 Wiley Periodicals, Inc.
Figures
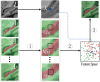


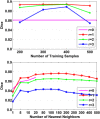
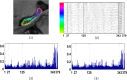
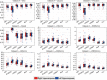
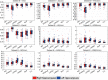
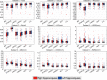



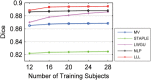

References
-
- Aljabar P, Heckeman R, Hammers A, Hajnal JV, Rueckert D (2007): Classifier selection strategies for label fusion using large atlas databases. Med Image Comput Comput Assist Interv 4791:523–531. - PubMed
-
- Aljabar P, Heckemann RA, Hammers A, Hajnal JV, Rueckert D (2009): Multi‐atlas based segmentation of brain images: Atlas selection and its effect on accuracy. Neuroimage 46:726–738. - PubMed
-
- Artaechevarria X, Munoz‐Barrutia A, Ortiz‐de‐Solorzano C (2008): Effleient classifier generation and weighted voting for aflas‐based segmentation: Two small steps faster and closer to the combination oracle. SPIE Med Imag 2008:6914.
-
- Artaechevarria X, Munoz‐Barrutia A, Ortiz‐de‐Solorzano C (2009): Combination strategies in multi‐atlas image segmentation: Application to brain MR data. IEEE Trans Image Process 28:1266–1277. - PubMed
-
- Ashburner J, Friston KJ (2005): Unified segmentation. Neuroimage 26:839–851. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

