Human cytomegalovirus UL50 and UL53 recruit viral protein kinase UL97, not protein kinase C, for disruption of nuclear lamina and nuclear egress in infected cells
- PMID: 24155370
- PMCID: PMC3911691
- DOI: 10.1128/JVI.02358-13
Human cytomegalovirus UL50 and UL53 recruit viral protein kinase UL97, not protein kinase C, for disruption of nuclear lamina and nuclear egress in infected cells
Abstract
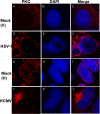
Herpesvirus nucleocapsids traverse the nuclear envelope into the cytoplasm in a process called nuclear egress that includes disruption of the nuclear lamina. In several herpesviruses, a key player in nuclear egress is a complex of two proteins, whose homologs in human cytomegalovirus (HCMV) are UL50 and UL53. However, their roles in nuclear egress during HCMV infection have not been shown. Based largely on transfection studies, UL50 and UL53 have been proposed to facilitate disruption of the nuclear lamina by recruiting cellular protein kinase C (PKC), as occurs with certain other herpesviruses, and/or the viral protein kinase UL97 to phosphorylate lamins. To investigate these issues during HCMV infection, we generated viral mutants null for UL50 or UL53. Correlative light electron microscopic analysis of null mutant-infected cells showed the presence of intranuclear nucleocapsids and the absence of cytoplasmic nucleocapsids. Confocal immunofluorescence microscopy revealed that UL50 and UL53 are required for disruption of the nuclear lamina. A subpopulation of UL97 colocalized with the nuclear rim, and this was dependent on UL50 and, to a lesser extent, UL53. However, PKC was not recruited to the nuclear rim, and its localization was not affected by the absence of UL50 or UL53. Immunoprecipitation from cells infected with HCMV expressing tagged UL53 detected UL97 but not PKC. In summary, HCMV UL50 and UL53 are required for nuclear egress and disruption of nuclear lamina during HCMV infection, and they recruit UL97, not PKC, for these processes. Thus, despite the strong conservation of herpesvirus nuclear egress complexes, a key function can differ among them.
Figures







Similar articles
-
Human cytomegalovirus UL97 phosphorylates the viral nuclear egress complex.J Virol. 2015 Jan;89(1):523-34. doi: 10.1128/JVI.02426-14. Epub 2014 Oct 22. J Virol. 2015. PMID: 25339763 Free PMC article.
-
The absence of p53 during Human Cytomegalovirus infection leads to decreased UL53 expression, disrupting UL50 localization to the inner nuclear membrane, and thereby inhibiting capsid nuclear egress.Virology. 2016 Oct;497:262-278. doi: 10.1016/j.virol.2016.07.020. Epub 2016 Aug 4. Virology. 2016. PMID: 27498409 Free PMC article.
-
Structure of a herpesvirus nuclear egress complex subunit reveals an interaction groove that is essential for viral replication.Proc Natl Acad Sci U S A. 2015 Jul 21;112(29):9010-5. doi: 10.1073/pnas.1511140112. Epub 2015 Jul 6. Proc Natl Acad Sci U S A. 2015. PMID: 26150520 Free PMC article.
-
Regulatory roles of protein kinases in cytomegalovirus replication.Adv Virus Res. 2011;80:69-101. doi: 10.1016/B978-0-12-385987-7.00004-X. Adv Virus Res. 2011. PMID: 21762822 Review.
-
Function of human cytomegalovirus UL97 kinase in viral infection and its inhibition by maribavir.Rev Med Virol. 2009 Jul;19(4):215-29. doi: 10.1002/rmv.615. Rev Med Virol. 2009. PMID: 19434630 Free PMC article. Review.
Cited by
-
Discovery of a Coregulatory Interaction between Kaposi's Sarcoma-Associated Herpesvirus ORF45 and the Viral Protein Kinase ORF36.J Virol. 2016 Jun 10;90(13):5953-5964. doi: 10.1128/JVI.00516-16. Print 2016 Jul 1. J Virol. 2016. PMID: 27099309 Free PMC article.
-
Functional Identification and Characterization of the Nuclear Egress Complex of a Gammaherpesvirus.J Virol. 2019 Nov 26;93(24):e01422-19. doi: 10.1128/JVI.01422-19. Print 2019 Dec 15. J Virol. 2019. PMID: 31554685 Free PMC article.
-
Nuclear Cytoskeleton in Virus Infection.Int J Mol Sci. 2022 Jan 5;23(1):578. doi: 10.3390/ijms23010578. Int J Mol Sci. 2022. PMID: 35009004 Free PMC article. Review.
-
Human cytomegalovirus pUL97 kinase induces global changes in the infected cell phosphoproteome.Proteomics. 2015 Jun;15(12):2006-22. doi: 10.1002/pmic.201400607. Epub 2015 May 12. Proteomics. 2015. PMID: 25867546 Free PMC article.
-
High-throughput analysis of human cytomegalovirus genome diversity highlights the widespread occurrence of gene-disrupting mutations and pervasive recombination.J Virol. 2015 Aug 1;89(15):7673-7695. doi: 10.1128/JVI.00578-15. Epub 2015 May 13. J Virol. 2015. PMID: 25972543 Free PMC article.
References
-
- Fuchs W, Klupp BG, Granzow H, Osterrieder N, Mettenleiter TC. 2002. The interacting UL31 and UL34 gene products of pseudorabies virus are involved in egress from the host-cell nucleus and represent components of primary enveloped but not mature virions. J. Virol. 76:364–378. 10.1128/JVI.76.1.364-378.2002 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

