Sequence signatures extracted from proximal promoters can be used to predict distal enhancers
- PMID: 24156763
- PMCID: PMC3983659
- DOI: 10.1186/gb-2013-14-10-r117
Sequence signatures extracted from proximal promoters can be used to predict distal enhancers
Abstract
Background: Gene expression is controlled by proximal promoters and distal regulatory elements such as enhancers. While the activity of some promoters can be invariant across tissues, enhancers tend to be highly tissue-specific.
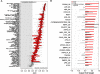
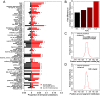
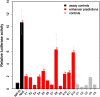
Results: We compiled sets of tissue-specific promoters based on gene expression profiles of 79 human tissues and cell types. Putative transcription factor binding sites within each set of sequences were used to train a support vector machine classifier capable of distinguishing tissue-specific promoters from control sequences. We obtained reliable classifiers for 92% of the tissues, with an area under the receiver operating characteristic curve between 60% (for subthalamic nucleus promoters) and 98% (for heart promoters). We next used these classifiers to identify tissue-specific enhancers, scanning distal non-coding sequences in the loci of the 200 most highly and lowly expressed genes. Thirty percent of reliable classifiers produced consistent enhancer predictions, with significantly higher densities in the loci of the most highly expressed compared to lowly expressed genes. Liver enhancer predictions were assessed in vivo using the hydrodynamic tail vein injection assay. Fifty-eight percent of the predictions yielded significant enhancer activity in the mouse liver, whereas a control set of five sequences was completely negative.
Conclusions: We conclude that promoters of tissue-specific genes often contain unambiguous tissue-specific signatures that can be learned and used for the de novo prediction of enhancers.
Figures
Similar articles
-
Sequence Characteristics Distinguish Transcribed Enhancers from Promoters and Predict Their Breadth of Activity.Genetics. 2019 Apr;211(4):1205-1217. doi: 10.1534/genetics.118.301895. Epub 2019 Jan 29. Genetics. 2019. PMID: 30696717 Free PMC article.
-
Taking promoters out of enhancers in sequence based predictions of tissue-specific mammalian enhancers.BMC Med Genomics. 2017 May 24;10(Suppl 1):34. doi: 10.1186/s12920-017-0264-3. BMC Med Genomics. 2017. PMID: 28589862 Free PMC article.
-
Integrating diverse datasets improves developmental enhancer prediction.PLoS Comput Biol. 2014 Jun 26;10(6):e1003677. doi: 10.1371/journal.pcbi.1003677. eCollection 2014 Jun. PLoS Comput Biol. 2014. PMID: 24967590 Free PMC article.
-
Evaluating Enhancer Function and Transcription.Annu Rev Biochem. 2020 Jun 20;89:213-234. doi: 10.1146/annurev-biochem-011420-095916. Epub 2020 Mar 20. Annu Rev Biochem. 2020. PMID: 32197056 Review.
-
Role of non-coding RNA transcription around gene regulatory elements in transcription factor recruitment.RNA Biol. 2017 Jan 2;14(1):1-5. doi: 10.1080/15476286.2016.1248020. Epub 2016 Oct 20. RNA Biol. 2017. PMID: 27763805 Free PMC article. Review.
Cited by
-
Novel principles of gamma-retroviral insertional transcription activation in murine leukemia virus-induced end-stage tumors.Retrovirology. 2014 May 19;11:36. doi: 10.1186/1742-4690-11-36. Retrovirology. 2014. PMID: 24886479 Free PMC article.
-
Comparative systeomics to elucidate physiological differences between CHO and SP2/0 cell lines.Sci Rep. 2022 Feb 28;12(1):3280. doi: 10.1038/s41598-022-06886-1. Sci Rep. 2022. PMID: 35228567 Free PMC article.
-
In silico identification of enhancers on the basis of a combination of transcription factor binding motif occurrences.Sci Rep. 2016 Sep 1;6:32476. doi: 10.1038/srep32476. Sci Rep. 2016. PMID: 27582178 Free PMC article.
-
cuRRBS: simple and robust evaluation of enzyme combinations for reduced representation approaches.Nucleic Acids Res. 2017 Nov 16;45(20):11559-11569. doi: 10.1093/nar/gkx814. Nucleic Acids Res. 2017. PMID: 29036576 Free PMC article.
-
A premature stop codon in BrFLC2 transcript results in early flowering in oilseed-type Brassica rapa plants.Plant Mol Biol. 2022 Feb;108(3):241-255. doi: 10.1007/s11103-021-01231-y. Epub 2022 Jan 22. Plant Mol Biol. 2022. PMID: 35064421
References
-
- Sandelin A, Carninci P, Lenhard B, Ponjavic J, Hayashizaki Y, Hume DA. Mammalian RNA polymerase II core promoters: insights from genome-wide studies. Nat Rev Genet. 2007;14:424–436. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P30 DK026743/DK/NIDDK NIH HHS/United States
- 1R01NS079231/NS/NINDS NIH HHS/United States
- CAPMC/ CIHR/Canada
- R01 HD059862/HD/NICHD NIH HHS/United States
- GM61390/GM/NIGMS NIH HHS/United States
- U01 GM061390/GM/NIGMS NIH HHS/United States
- ImNIH/Intramural NIH HHS/United States
- R01 DK090382/DK/NIDDK NIH HHS/United States
- T32 GM007175/GM/NIGMS NIH HHS/United States
- 1R01HG005058/HG/NHGRI NIH HHS/United States
- 1R01HG006768/HG/NHGRI NIH HHS/United States
- 1R01DK090382/DK/NIDDK NIH HHS/United States
- R01 NS079231/NS/NINDS NIH HHS/United States
- U19 GM061390/GM/NIGMS NIH HHS/United States
- R01HD059862/HD/NICHD NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources

