The hammerhead ribozyme: structure, catalysis, and gene regulation
- PMID: 24156940
- PMCID: PMC4008931
- DOI: 10.1016/B978-0-12-381286-5.00001-9
The hammerhead ribozyme: structure, catalysis, and gene regulation
Abstract
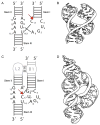
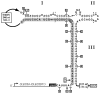
The hammerhead ribozyme has long been considered a prototype for understanding RNA catalysis, but discrepancies between the earlier crystal structures of a minimal hammerhead self-cleaving motif and various biochemical investigations frustrated attempt to understand hammerhead ribozyme catalysis in terms of structure. With the discovery that a tertiary contact distal from the ribozyme's active site greatly enhances its catalytic prowess, and the emergence of new corresponding crystal structures of full-length hammerhead ribozymes, a unified understanding of catalysis in terms of the structure is now possible. A mechanism in which the invariant residue G12 functions as a general base, and the 2'-OH moiety of the invariant G8, itself forming a tertiary base pair with the invariant C3, is the general acid, appears consistent with both the crystal structure and biochemical experimental results. Originally discovered in the context of plant satellite RNA viruses, the hammerhead more recently has been found embedded in the 3'-untranslated region of mature mammalian mRNAs, suggesting additional biological roles in genetic regulation.
Keywords: Catalysis; Gene regulation; Hammerhead ribozyme; RNA; Riboswitch.
© 2013 Elsevier Inc. All rights reserved.
Figures
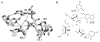
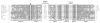
Similar articles
-
Structural Simplicity and Mechanistic Complexity in the Hammerhead Ribozyme.Prog Mol Biol Transl Sci. 2018;159:177-202. doi: 10.1016/bs.pmbts.2018.07.006. Epub 2018 Sep 17. Prog Mol Biol Transl Sci. 2018. PMID: 30340787 Review.
-
Structural and catalytic effects of an invariant purine substitution in the hammerhead ribozyme: implications for the mechanism of acid-base catalysis.Acta Crystallogr D Biol Crystallogr. 2014 Sep;70(Pt 9):2256-63. doi: 10.1107/S1399004714010608. Epub 2014 Aug 23. Acta Crystallogr D Biol Crystallogr. 2014. PMID: 25195740 Free PMC article.
-
Capturing hammerhead ribozyme structures in action by modulating general base catalysis.PLoS Biol. 2008 Sep 30;6(9):e234. doi: 10.1371/journal.pbio.0060234. PLoS Biol. 2008. PMID: 18834200 Free PMC article.
-
Probing general base catalysis in the hammerhead ribozyme.J Am Chem Soc. 2008 Nov 19;130(46):15467-75. doi: 10.1021/ja804496z. Epub 2008 Oct 25. J Am Chem Soc. 2008. PMID: 18950173
-
Catalytic strategies of self-cleaving ribozymes.Acc Chem Res. 2008 Aug;41(8):1027-35. doi: 10.1021/ar800050c. Epub 2008 Jul 25. Acc Chem Res. 2008. PMID: 18652494 Review.
Cited by
-
A Newly Identified Peripheral Duplex Anchors and Stabilizes the MALAT1 Triplex.Biochemistry. 2024 Sep 17;63(18):2280-2292. doi: 10.1021/acs.biochem.4c00156. Epub 2024 Aug 27. Biochemistry. 2024. PMID: 39190685 Free PMC article.
-
Rational design of aptazyme riboswitches for efficient control of gene expression in mammalian cells.Elife. 2016 Nov 2;5:e18858. doi: 10.7554/eLife.18858. Elife. 2016. PMID: 27805569 Free PMC article.
-
A single-vector intersectional AAV strategy for interrogating cellular diversity and brain function.Nat Neurosci. 2024 Jul;27(7):1400-1410. doi: 10.1038/s41593-024-01659-7. Epub 2024 May 27. Nat Neurosci. 2024. PMID: 38802592
-
Nucleotide Selectivity in Abiotic RNA Polymerization Reactions.Orig Life Evol Biosph. 2017 Sep;47(3):305-321. doi: 10.1007/s11084-017-9532-7. Epub 2017 Feb 3. Orig Life Evol Biosph. 2017. PMID: 28160163 Free PMC article.
-
Strategies for detecting aminoacylation and aminoacyl-tRNA editing in vitro and in cells.Isr J Chem. 2024 Sep;64(8-9):e202400009. doi: 10.1002/ijch.202400009. Epub 2024 May 6. Isr J Chem. 2024. PMID: 40066018 Free PMC article.
References
-
- Kruger K, Grabowski PJ, Zaug AJ, et al. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell. 1982;31:147–157. - PubMed
-
- Guerrier-Takada C, Gardiner K, Marsh T, Pace N, Altman S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell. 1983;35:849–857. - PubMed
-
- Prody GA, Bakos JT, Buzayan JM, et al. Autolytic processing of dimeric plant virus satellite RNA. Science. 1986;231:1577–1580. - PubMed
-
- Noller HF, Hoffarth V, Zimniak L. Unusual resistance of peptidyl transferase to protein extraction procedures. Science. 1992;256:1416–1419. - PubMed
-
- Nissen P, Hansen J, Ban N, et al. The structural basis of ribosome activity in peptide bond synthesis. Science. 2000;289:920–930. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

