ETV1 positively regulates transcription of tumor suppressor ARF
- PMID: 24157551
- PMCID: PMC3912040
- DOI: 10.4161/cbt.26883
ETV1 positively regulates transcription of tumor suppressor ARF
Abstract
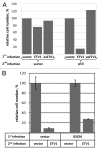
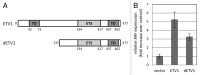
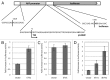
ETV1 (ETS variant 1) is a transcription factor from the ETS family and an oncogene in several types of human malignancies. Paradoxically, a predicted inactivating mutation in ETV1 was previously found in a clone of HT1080 cells with reduced activity of p53. We report that elevated expression of ETV1 makes p53-null tumor cells hypersensitive to restoration of said tumor suppressor. Furthermore, elevated levels of either wild-type ETV1 or its truncated derivative, dETV1, which mimics the product of an oncogenic rearrangement in certain tumors, results in increased expression of mRNA for p14ARF, a known activator of p53. Accordingly, expression of a luciferase reporter, which is driven by a putative ARF promoter, was elevated by concomitant expression of either ETV1 or dETV1. Our observations point to yet another example of a tumor suppressor gene being activated by a potentially oncogenic signal. A better understanding of the mechanisms that allow a cell to bypass such safeguards is needed in order to predict and prevent the development of an oncogene-tolerant state during cancer evolution.
Keywords: ETV1; oncogenic transformation; p14ARF; p53; tumor suppression.
Figures
Similar articles
-
Azidothymidine and cisplatin increase p14ARF expression in OVCAR-3 ovarian cancer cell line.Toxicol Appl Pharmacol. 2006 Oct 1;216(1):89-97. doi: 10.1016/j.taap.2006.04.015. Epub 2006 May 19. Toxicol Appl Pharmacol. 2006. PMID: 16797627
-
p19(ARF) is dispensable for oncogenic stress-induced p53-mediated apoptosis and tumor suppression in vivo.Mol Cell Biol. 2002 Jan;22(1):370-7. doi: 10.1128/MCB.22.1.370-377.2002. Mol Cell Biol. 2002. PMID: 11739748 Free PMC article.
-
ARF directly binds DP1: interaction with DP1 coincides with the G1 arrest function of ARF.Mol Cell Biol. 2005 Sep;25(18):8024-36. doi: 10.1128/MCB.25.18.8024-8036.2005. Mol Cell Biol. 2005. PMID: 16135794 Free PMC article.
-
Regulation of the p14ARF-Mdm2-p53 pathway: an overview in breast cancer.Exp Mol Pathol. 2006 Oct;81(2):115-22. doi: 10.1016/j.yexmp.2006.07.001. Epub 2006 Aug 17. Exp Mol Pathol. 2006. PMID: 16919268 Review.
-
E2F3-a novel repressor of the ARF/p53 pathway.Dev Cell. 2004 Jun;6(6):742-3. doi: 10.1016/j.devcel.2004.05.012. Dev Cell. 2004. PMID: 15177020 Review.
Cited by
-
miR-17 regulates melanoma cell motility by inhibiting the translation of ETV1.Oncotarget. 2015 Aug 7;6(22):19006-16. doi: 10.18632/oncotarget.4147. Oncotarget. 2015. PMID: 26158900 Free PMC article.
-
SUMO modified ETV1 promotes M2-polarized tumor-associated macrophage infiltration and cancer progression by facilitating CCL2 transcription in esophageal squamous cell carcinoma cells.Cancer Immunol Immunother. 2025 Feb 1;74(3):87. doi: 10.1007/s00262-024-03914-z. Cancer Immunol Immunother. 2025. PMID: 39891717 Free PMC article.
-
MicroRNA analysis suggests an additional level of feedback regulation in the NF-κB signaling cascade.Oncotarget. 2015 Jul 10;6(19):17097-106. doi: 10.18632/oncotarget.4005. Oncotarget. 2015. PMID: 26020802 Free PMC article.
-
Molecular Mechanism of β-Catenin Signaling Pathway Inactivation in ETV1-Positive Prostate Cancers.J Pharm Sci Pharmacol. 2015 Sep;2(3):208-216. doi: 10.1166/jpsp.2015.1069. J Pharm Sci Pharmacol. 2015. PMID: 28497076 Free PMC article.
References
-
- Galang CK, García-Ramírez J, Solski PA, Westwick JK, Der CJ, Neznanov NN, Oshima RG, Hauser CA. Oncogenic Neu/ErbB-2 increases ets, AP-1, and NF-kappaB-dependent gene expression, and inhibiting ets activation blocks Neu-mediated cellular transformation. J Biol Chem. 1996;271:7992–8. doi: 10.1074/jbc.271.14.7992. - DOI - PubMed
-
- Seth A, Papas TS. The c-ets-1 proto-oncogene has oncogenic activity and is positively autoregulated. Oncogene. 1990;5:1761–7. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
