Promoter decoding of transcription factor dynamics involves a trade-off between noise and control of gene expression
- PMID: 24189399
- PMCID: PMC4039373
- DOI: 10.1038/msb.2013.56
Promoter decoding of transcription factor dynamics involves a trade-off between noise and control of gene expression
Abstract
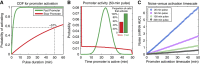
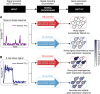
Numerous transcription factors (TFs) encode information about upstream signals in the dynamics of their activation, but how downstream genes decode these dynamics remains poorly understood. Using microfluidics to control the nucleocytoplasmic translocation dynamics of the budding yeast TF Msn2, we elucidate the principles that govern how different promoters convert dynamical Msn2 input into gene expression output in single cells. Combining modeling and experiments, we classify promoters according to their signal-processing behavior and reveal that multiple, distinct gene expression programs can be encoded in the dynamics of Msn2. We show that both oscillatory TF dynamics and slow promoter kinetics lead to higher noise in gene expression. Furthermore, we show that the promoter activation timescale is related to nucleosome remodeling. Our findings imply a fundamental trade-off: although the cell can exploit different promoter classes to differentially control gene expression using TF dynamics, gene expression noise fundamentally limits how much information can be encoded in the dynamics of a single TF and reliably decoded by promoters.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures






Comment in
-
Promoter decoding of transcription factor dynamics.Mol Syst Biol. 2013 Nov 5;9:703. doi: 10.1038/msb.2013.63. Mol Syst Biol. 2013. PMID: 24189398 Free PMC article. No abstract available.
References
-
- Balaban NQ, Merrin J, Chait R, Kowalik L, Leibler S (2004) Bacterial persistence as a phenotypic switch. Science 305: 1622–1625 - PubMed
-
- Bar-Even A, Paulsson J, Maheshri N, Carmi M, O'Shea E, Pilpel Y, Barkai N (2006) Noise in protein expression scales with natural protein abundance. Nat Genet 38: 636–643 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

