Murine neural stem cells model Hunter disease in vitro: glial cell-mediated neurodegeneration as a possible mechanism involved
- PMID: 24201805
- PMCID: PMC3847312
- DOI: 10.1038/cddis.2013.430
Murine neural stem cells model Hunter disease in vitro: glial cell-mediated neurodegeneration as a possible mechanism involved
Abstract
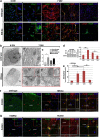
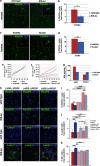
Mucopolysaccharidosis type II (MPSII or Hunter Syndrome) is a lysosomal storage disorder caused by the deficit of iduronate 2-sulfatase (IDS) activity and characterized by progressive systemic and neurological impairment. As the early mechanisms leading to neuronal degeneration remain elusive, we chose to examine the properties of neural stem cells (NSCs) isolated from an animal model of the disease in order to evaluate whether their neurogenic potential could be used to recapitulate the early phases of neurogenesis in the brain of Hunter disease patients. Experiments here reported show that NSCs derived from the subventricular zone (SVZ) of early symptomatic IDS-knockout (IDS-ko) mouse retained self-renewal capacity in vitro, but differentiated earlier than wild-type (wt) cells, displaying an evident lysosomal aggregation in oligodendroglial and astroglial cells. Consistently, the SVZ of IDS-ko mice appeared similar to the wt SVZ, whereas the cortex and striatum presented a disorganized neuronal pattern together with a significant increase of glial apoptotic cells, suggesting that glial degeneration likely precedes neuronal demise. Interestingly, a very similar pattern was observed in the brain cortex of a Hunter patient. These observations both in vitro, in our model, and in vivo suggest that IDS deficit seems to affect the late phases of neurogenesis and/or the survival of mature cells rather than NSC self-renewal. In particular, platelet-derived growth factor receptor-α-positive (PDGFR-α+) glial progenitors appeared reduced in both the IDS-ko NSCs and in the IDS-ko mouse and human Hunter brains, compared with the respective healthy controls. Treatment of mutant NSCs with IDS or PDGF throughout differentiation was able to increase the number of PDGFR-α+ cells and to reduce that of apoptotic cells to levels comparable to wt. This evidence supports IDS-ko NSCs as a reliable in vitro model of the disease, and suggests the rescue of PDGFR-α+ glial cells as a therapeutic strategy to prevent neuronal degeneration.
Figures



Similar articles
-
Neural Stem Cells from Shank3-ko Mouse Model Autism Spectrum Disorders.Mol Neurobiol. 2020 Mar;57(3):1502-1515. doi: 10.1007/s12035-019-01811-6. Epub 2019 Nov 26. Mol Neurobiol. 2020. PMID: 31773410
-
Glial degeneration with oxidative damage drives neuronal demise in MPSII disease.Cell Death Dis. 2016 Aug 11;7(8):e2331. doi: 10.1038/cddis.2016.231. Cell Death Dis. 2016. PMID: 27512952 Free PMC article.
-
Neural cells generated from human induced pluripotent stem cells as a model of CNS involvement in mucopolysaccharidosis type II.J Inherit Metab Dis. 2018 Mar;41(2):221-229. doi: 10.1007/s10545-017-0108-5. Epub 2017 Nov 22. J Inherit Metab Dis. 2018. PMID: 29168031
-
Current and Future Treatment of Mucopolysaccharidosis (MPS) Type II: Is Brain-Targeted Stem Cell Gene Therapy the Solution for This Devastating Disorder?Int J Mol Sci. 2022 Apr 27;23(9):4854. doi: 10.3390/ijms23094854. Int J Mol Sci. 2022. PMID: 35563245 Free PMC article. Review.
-
A molecular genetics view on Mucopolysaccharidosis Type II.Mutat Res Rev Mutat Res. 2021 Jul-Dec;788:108392. doi: 10.1016/j.mrrev.2021.108392. Epub 2021 Aug 8. Mutat Res Rev Mutat Res. 2021. PMID: 34893157 Review.
Cited by
-
Neural Stem Cells from Shank3-ko Mouse Model Autism Spectrum Disorders.Mol Neurobiol. 2020 Mar;57(3):1502-1515. doi: 10.1007/s12035-019-01811-6. Epub 2019 Nov 26. Mol Neurobiol. 2020. PMID: 31773410
-
Dose-dependent effects of a brain-penetrating iduronate-2-sulfatase on neurobehavioral impairments in mucopolysaccharidosis II mice.Mol Ther Methods Clin Dev. 2022 May 10;25:534-544. doi: 10.1016/j.omtm.2022.05.002. eCollection 2022 Jun 9. Mol Ther Methods Clin Dev. 2022. PMID: 35662814 Free PMC article.
-
Mucopolysaccharidosis Type II: One Hundred Years of Research, Diagnosis, and Treatment.Int J Mol Sci. 2020 Feb 13;21(4):1258. doi: 10.3390/ijms21041258. Int J Mol Sci. 2020. PMID: 32070051 Free PMC article. Review.
-
Pathogenic Roles of Heparan Sulfate and Its Use as a Biomarker in Mucopolysaccharidoses.Int J Mol Sci. 2022 Oct 3;23(19):11724. doi: 10.3390/ijms231911724. Int J Mol Sci. 2022. PMID: 36233030 Free PMC article. Review.
-
Iduronate-2-sulfatase interactome: validation by yeast two-hybrid assay.Heliyon. 2022 Mar 1;8(3):e09031. doi: 10.1016/j.heliyon.2022.e09031. eCollection 2022 Mar. Heliyon. 2022. PMID: 35284671 Free PMC article.
References
-
- Muenzer J. Overview of the mucopolysaccharidoses. Rheumatology (Oxford) 2011;50:v4–v12. - PubMed
-
- Muenzer J, Beck M, Eng CM, Giugliani R, Harmatz P, Martin R, et al. Long-term, open-labeled extension study of idursulfase in the treatment of Hunter syndrome. Genet Med. 2011;13:95–101. - PubMed
-
- Muenzer J, Beck M, Giugliani R, Suzuki Y, Tylki-Szymanska A, Valayannopoulos V, et al. Idursulfase treatment of Hunter syndrome in children younger than 6 years: results from the Hunter Outcome Survey. Genet Med. 2011;13:102–109. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

