Genome-wide high-resolution mapping of UV-induced mitotic recombination events in Saccharomyces cerevisiae
- PMID: 24204306
- PMCID: PMC3814309
- DOI: 10.1371/journal.pgen.1003894
Genome-wide high-resolution mapping of UV-induced mitotic recombination events in Saccharomyces cerevisiae
Abstract
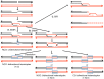
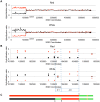
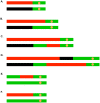
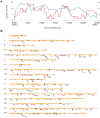
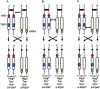
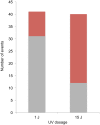
In the yeast Saccharomyces cerevisiae and most other eukaryotes, mitotic recombination is important for the repair of double-stranded DNA breaks (DSBs). Mitotic recombination between homologous chromosomes can result in loss of heterozygosity (LOH). In this study, LOH events induced by ultraviolet (UV) light are mapped throughout the genome to a resolution of about 1 kb using single-nucleotide polymorphism (SNP) microarrays. UV doses that have little effect on the viability of diploid cells stimulate crossovers more than 1000-fold in wild-type cells. In addition, UV stimulates recombination in G1-synchronized cells about 10-fold more efficiently than in G2-synchronized cells. Importantly, at high doses of UV, most conversion events reflect the repair of two sister chromatids that are broken at approximately the same position whereas at low doses, most conversion events reflect the repair of a single broken chromatid. Genome-wide mapping of about 380 unselected crossovers, break-induced replication (BIR) events, and gene conversions shows that UV-induced recombination events occur throughout the genome without pronounced hotspots, although the ribosomal RNA gene cluster has a significantly lower frequency of crossovers.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Similar articles
-
High-resolution genome-wide analysis of irradiated (UV and γ-rays) diploid yeast cells reveals a high frequency of genomic loss of heterozygosity (LOH) events.Genetics. 2012 Apr;190(4):1267-84. doi: 10.1534/genetics.111.137927. Epub 2012 Jan 20. Genetics. 2012. PMID: 22267500 Free PMC article.
-
High-Resolution Mapping of Homologous Recombination Events in rad3 Hyper-Recombination Mutants in Yeast.PLoS Genet. 2016 Mar 11;12(3):e1005938. doi: 10.1371/journal.pgen.1005938. eCollection 2016 Mar. PLoS Genet. 2016. PMID: 26968037 Free PMC article.
-
High-resolution mapping of spontaneous mitotic recombination hotspots on the 1.1 Mb arm of yeast chromosome IV.PLoS Genet. 2013 Apr;9(4):e1003434. doi: 10.1371/journal.pgen.1003434. Epub 2013 Apr 4. PLoS Genet. 2013. PMID: 23593029 Free PMC article.
-
Mitotic recombination in Saccharomyces cerevisiae.Curr Genet. 2003 Jan;42(4):185-98. doi: 10.1007/s00294-002-0346-3. Epub 2002 Nov 29. Curr Genet. 2003. PMID: 12589470 Review.
-
Double-strand break-induced recombination in eukaryotes.Prog Nucleic Acid Res Mol Biol. 1998;58:263-99. doi: 10.1016/s0079-6603(08)60039-2. Prog Nucleic Acid Res Mol Biol. 1998. PMID: 9308369 Review.
Cited by
-
Loss of Heterozygosity and Base Mutation Rates Vary Among Saccharomyces cerevisiae Hybrid Strains.G3 (Bethesda). 2020 Sep 2;10(9):3309-3319. doi: 10.1534/g3.120.401551. G3 (Bethesda). 2020. PMID: 32727920 Free PMC article.
-
Heterozygous mutations cause genetic instability in a yeast model of cancer evolution.Nature. 2019 Feb;566(7743):275-278. doi: 10.1038/s41586-019-0887-y. Epub 2019 Jan 30. Nature. 2019. PMID: 30700905 Free PMC article.
-
Effects of Temperature on the Meiotic Recombination Landscape of the Yeast Saccharomyces cerevisiae.mBio. 2017 Dec 19;8(6):e02099-17. doi: 10.1128/mBio.02099-17. mBio. 2017. PMID: 29259092 Free PMC article.
-
Genome Dynamics of Hybrid Saccharomyces cerevisiae During Vegetative and Meiotic Divisions.G3 (Bethesda). 2017 Nov 6;7(11):3669-3679. doi: 10.1534/g3.117.1135. G3 (Bethesda). 2017. PMID: 28916648 Free PMC article.
-
Mechanisms and regulation of mitotic recombination in Saccharomyces cerevisiae.Genetics. 2014 Nov;198(3):795-835. doi: 10.1534/genetics.114.166140. Genetics. 2014. PMID: 25381364 Free PMC article. Review.
References
-
- Friedberg EC, Walker GC, Siede W, Schultz RA (2006) DNA repair and mutagenesis. Washington, D.C.: ASM Press.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

