Microarray analysis of tomato's early and late wound response reveals new regulatory targets for Leucine aminopeptidase A
- PMID: 24205013
- PMCID: PMC3812031
- DOI: 10.1371/journal.pone.0077889
Microarray analysis of tomato's early and late wound response reveals new regulatory targets for Leucine aminopeptidase A
Abstract
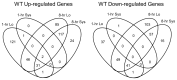
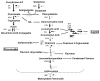
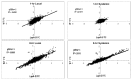
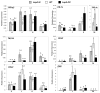
Wounding due to mechanical injury or insect feeding causes a wide array of damage to plant cells including cell disruption, desiccation, metabolite oxidation, and disruption of primary metabolism. In response, plants regulate a variety of genes and metabolic pathways to cope with injury. Tomato (Solanum lycopersicum) is a model for wound signaling but few studies have examined the comprehensive gene expression profiles in response to injury. A cross-species microarray approach using the TIGR potato 10-K cDNA array was analyzed for large-scale temporal (early and late) and spatial (locally and systemically) responses to mechanical wounding in tomato leaves. These analyses demonstrated that tomato regulates many primary and secondary metabolic pathways and this regulation is dependent on both timing and location. To determine if LAP-A, a known modulator of wound signaling, influences gene expression beyond the core of late wound-response genes, changes in RNAs from healthy and wounded Leucine aminopeptidase A-silenced (LapA-SI) and wild-type (WT) leaves were examined. While most of the changes in gene expression after wounding in LapA-SI leaves were similar to WT, overall responses were delayed in the LapA-SI leaves. Moreover, two pathogenesis-related 1 (PR-1c and PR-1a2) and two dehydrin (TAS14 and Dhn3) genes were negatively regulated by LAP-A. Collectively, this study has shown that tomato wound responses are complex and that LAP-A's role in modulation of wound responses extends beyond the well described late-wound gene core.
Conflict of interest statement
Figures


Similar articles
-
The induction of tomato leucine aminopeptidase genes (LapA) after Pseudomonas syringae pv. tomato infection is primarily a wound response triggered by coronatine.Mol Plant Microbe Interact. 2001 Feb;14(2):214-24. doi: 10.1094/MPMI.2001.14.2.214. Mol Plant Microbe Interact. 2001. PMID: 11204785
-
Leucine aminopeptidase regulates defense and wound signaling in tomato downstream of jasmonic acid.Plant Cell. 2009 Apr;21(4):1239-51. doi: 10.1105/tpc.108.065029. Epub 2009 Apr 17. Plant Cell. 2009. PMID: 19376935 Free PMC article.
-
Leucine aminopeptidases: the ubiquity of LAP-N and the specificity of LAP-A.Planta. 2000 Mar;210(4):563-73. doi: 10.1007/s004250050045. Planta. 2000. PMID: 10787049
-
A -308 deletion of the tomato LAP promoters is able to direct flower-specific and MeJA-induced expression in transgenic plants.Plant Mol Biol. 1998 Mar;36(5):639-48. doi: 10.1023/a:1005980028203. Plant Mol Biol. 1998. PMID: 9526496
-
Overexpression, purification and biochemical characterization of the wound-induced leucine aminopeptidase of tomato.Eur J Biochem. 1999 Aug;263(3):726-35. doi: 10.1046/j.1432-1327.1999.00548.x. Eur J Biochem. 1999. PMID: 10469136
Cited by
-
Modifying Anthocyanins Biosynthesis in Tomato Hairy Roots: A Test Bed for Plant Resistance to Ionizing Radiation and Antioxidant Properties in Space.Front Plant Sci. 2022 Feb 24;13:830931. doi: 10.3389/fpls.2022.830931. eCollection 2022. Front Plant Sci. 2022. PMID: 35283922 Free PMC article.
-
Isolation and characterization of systemic acquired resistance marker gene PR1 and its promoter from Brassica juncea.3 Biotech. 2018 Jan;8(1):10. doi: 10.1007/s13205-017-1027-8. Epub 2017 Dec 11. 3 Biotech. 2018. PMID: 29259885 Free PMC article.
-
Mechanisms and ecological consequences of plant defence induction and suppression in herbivore communities.Ann Bot. 2015 Jun;115(7):1015-51. doi: 10.1093/aob/mcv054. Ann Bot. 2015. PMID: 26019168 Free PMC article. Review.
-
Simulated herbivory in chickpea causes rapid changes in defense pathways and hormonal transcription networks of JA/ethylene/GA/auxin within minutes of wounding.Sci Rep. 2017 Mar 16;7:44729. doi: 10.1038/srep44729. Sci Rep. 2017. PMID: 28300183 Free PMC article.
-
A rapid preparation procedure for laser microdissection-mediated harvest of plant tissues for gene expression analysis.Plant Methods. 2019 Aug 2;15:88. doi: 10.1186/s13007-019-0471-3. eCollection 2019. Plant Methods. 2019. PMID: 31388345 Free PMC article.
References
-
- Walling LL (2009) Adaptive defense responses to pathogens and insects. Adv Bot Res 51: 551-612. doi:10.1016/S0065-2296(09)51013-0. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

