ALDH-1 expression levels predict response or resistance to preoperative chemoradiation in resectable esophageal cancer patients
- PMID: 24210755
- PMCID: PMC3946849
- DOI: 10.1016/j.molonc.2013.10.007
ALDH-1 expression levels predict response or resistance to preoperative chemoradiation in resectable esophageal cancer patients
Abstract
Purpose: Operable thoracic esophageal/gastroesophageal junction carcinoma (EC) is often treated with chemoradiation and surgery but tumor responses are unpredictable and heterogeneous. We hypothesized that aldehyde dehydrogenase-1 (ALDH-1) could be associated with response.
Methods: The labeling indices (LIs) of ALDH-1 by immunohistochemistry in untreated tumor specimens were established in EC patients who had chemoradiation and surgery. Univariate logistic regression and 3-fold cross validation were carried out for the training (67% of patients) and validation (33%) sets. Non-clinical experiments in EC cells were performed to generate complimentary data.
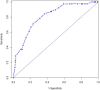
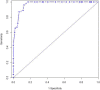
Results: Of 167 EC patients analyzed, 40 (24%) had a pathologic complete response (pathCR) and 27 (16%) had an extremely resistant (exCRTR) cancer. The median ALDH-1 LI was 0.2 (range, 0.01-0.85). There was a significant association between pathCR and low ALDH-1 LI (p ≤ 0.001; odds-ratio [OR] = 0.432). The 3-fold cross validation led to a concordance index (C-index) of 0.798 for the fitted model. There was a significant association between exCRTR and high ALDH-1 LI (p ≤ 0.001; OR = 3.782). The 3-fold cross validation led to the C-index of 0.960 for the fitted model. In several cell lines, higher ALDH-1 LIs correlated with resistant/aggressive phenotype. Cells with induced chemotherapy resistance upregulated ALDH-1 and resistance conferring genes (SOX9 and YAP1). Sorted ALDH-1+ cells were more resistant and had an aggressive phenotype in tumor spheres than ALDH-1- cells.
Conclusions: Our clinical and non-clinical data demonstrate that ALDH-1 LIs are predictive of response to therapy and further research could lead to individualized therapeutic strategies and novel therapeutic targets for EC patients.
Keywords: ALDH-1; Chemotherapy resistance; Esophageal carcinoma; Predictive markers; Prognosis; Radiation resistance.
Copyright © 2013 Federation of European Biochemical Societies. Published by Elsevier B.V. All rights reserved.
Figures



References
-
- Ajani, J.A. , Barthel, J.S. , Bentrem, D.J. , 2011. Esophageal and esophagogastric junction cancers. J. Natl. Compr. Canc Netw.. 9, 830–887. - PubMed
-
- Allum, W.H. , Stenning, S.P. , Bancewicz, J. , Clark, P.I. , Langley, R.E. , 2009. Long-term results of a randomized trial of surgery with or without preoperative chemotherapy in esophageal cancer. J. Clin. Oncol.. 27, 5062–5067. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

