Interferon-dependent IL-10 production by Tregs limits tumor Th17 inflammation
- PMID: 24216477
- PMCID: PMC3809773
- DOI: 10.1172/JCI65180
Interferon-dependent IL-10 production by Tregs limits tumor Th17 inflammation
Abstract
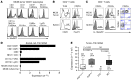
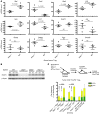
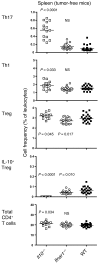
The capacity of IL-10 and Tregs in the inflammatory tumor microenvironment to impair anticancer Th1 immunity makes them attractive targets for cancer immunotherapy. IL-10 and Tregs also suppress Th17 activity, which is associated with poor prognosis in several cancers. However, previous studies have overlooked their potential contribution to the regulation of pathogenic cancer-associated inflammation. In this study, we investigated the origin and function of IL-10–producing cells in the tumor microenvironment using transplantable tumor models in mice. The majority of tumor-associated IL-10 was produced by an activated Treg population. IL-10 production by Tregs was required to restrain Th17-type inflammation. Accumulation of activated IL-10+ Tregs in the tumor required type I IFN signaling but not inflammatory signaling pathways that depend on TLR adapter protein MyD88 or IL-12 family cytokines. IL-10 production limited Th17 cell numbers in both spleen and tumor. However, type I IFN was required to limit Th17 cells specifically in the tumor microenvironment, reflecting selective control of tumor-associated Tregs by type I IFN. Thus, the interplay of type I IFN, Tregs, and IL-10 is required to negatively regulate Th17 inflammation in the tumor microenvironment. Therapeutic interference of this network could therefore have the undesirable consequence of promoting Th17 inflammation and cancer growth.
Figures






References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

