Frataxin directly stimulates mitochondrial cysteine desulfurase by exposing substrate-binding sites, and a mutant Fe-S cluster scaffold protein with frataxin-bypassing ability acts similarly
- PMID: 24217246
- PMCID: PMC3873537
- DOI: 10.1074/jbc.M113.525857
Frataxin directly stimulates mitochondrial cysteine desulfurase by exposing substrate-binding sites, and a mutant Fe-S cluster scaffold protein with frataxin-bypassing ability acts similarly
Abstract
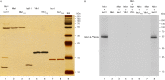
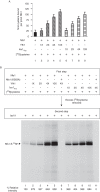
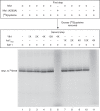
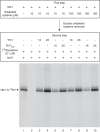
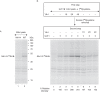
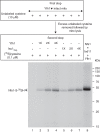
For iron-sulfur (Fe-S) cluster synthesis in mitochondria, the sulfur is derived from the amino acid cysteine by the cysteine desulfurase activity of Nfs1. The enzyme binds the substrate cysteine in the pyridoxal phosphate-containing site, and a persulfide is formed on the active site cysteine in a manner depending on the accessory protein Isd11. The persulfide is then transferred to the scaffold Isu, where it combines with iron to form the Fe-S cluster intermediate. Frataxin is implicated in the process, although it is unclear where and how, and deficiency causes Friedreich ataxia. Using purified proteins and isolated mitochondria, we show here that the yeast frataxin homolog (Yfh1) directly and specifically stimulates cysteine binding to Nfs1 by exposing substrate-binding sites. This novel function of frataxin does not require iron, Isu1, or Isd11. Once bound to Nfs1, the substrate cysteine is the source of the Nfs1 persulfide, but this step is independent of frataxin and strictly dependent on Isd11. Recently, a point mutation in Isu1 was found to bypass many frataxin functions. The data presented here show that the Isu1 suppressor mimics the frataxin effects on Nfs1, explaining the bypassing activity. We propose a regulatory mechanism for the Nfs1 persulfide-forming activity. Specifically, at least two separate conformational changes must occur in the enzyme for optimum activity as follows: one is mediated by frataxin interaction that exposes the "buried" substrate-binding sites, and the other is mediated by Isd11 interaction that brings the bound substrate cysteine and the active site cysteine in proximity for persulfide formation.
Keywords: Iron-Sulfur Protein; Metal Homeostasis; Metals; Mitochondria; Mitochondrial Metabolism; Pyridoxal Phosphate; Sulfur; Yeast.
Figures





References
-
- Lill R., Hoffmann B., Molik S., Pierik A. J., Rietzschel N., Stehling O., Uzarska M. A., Webert H., Wilbrecht C., Mühlenhoff U. (2012) The role of mitochondria in cellular iron-sulfur protein biogenesis and iron metabolism. Biochim. Biophys. Acta 1823, 1491–1508 - PubMed
-
- Campuzano V., Montermini L., Moltò M. D., Pianese L., Cossée M., Cavalcanti F., Monros E., Rodius F., Duclos F., Monticelli A., Zara F., Cañizares J., Koutnikova H., Bidichandani S. I., Gellera C., Brice A., Trouillas P., De Michele G., Filla A., De Frutos R., Palau F., Patel P. I., Di Donato S., Mandel J. L., Cocozza S., Koenig M., Pandolfo M. (1996) Friedreich's ataxia: autosomal recessive disease caused by an intronic GAA triplet repeat expansion. Science 271, 1423–1427 - PubMed
-
- Rötig A., de Lonlay P., Chretien D., Foury F., Koenig M., Sidi D., Munnich A., Rustin P. (1997) Aconitase and mitochondrial iron-sulfur protein deficiency in Friedreich ataxia. Nat. Genet. 17, 215–217 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

