Specificity in the actions of the UBR1 ubiquitin ligase in the degradation of nuclear receptors
- PMID: 24251101
- PMCID: PMC3821023
- DOI: 10.1016/j.fob.2013.09.003
Specificity in the actions of the UBR1 ubiquitin ligase in the degradation of nuclear receptors
Abstract
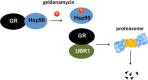
The UBR1 ubiquitin ligase promotes degradation of proteins via the N-end rule and by another mechanism that detects a misfolded conformation. Although UBR1 was shown recently to act on protein kinases whose misfolding was promoted by inhibition of Hsp90, it was unknown whether this ubiquitin ligase targeted other client types of the chaperone. We analyzed the role of UBR1 in the degradation of nuclear receptors that are classical clients of Hsp90. Our results showed that UBR1 deletion results in impaired degradation of the glucocorticoid receptor and the androgen receptor but not the estrogen receptor α. These findings demonstrate specificity in the actions of the UBR1 ubiquitin ligase in the degradation of Hsp90 clients in the presence of small molecule inhibitors that promote client misfolding.
Keywords: AR, androgen receptor; CHIP, C-terminal Hsp interacting protein; DMSO, dimethylsulfoxide; Degradation; ER, estrogen receptor; GA, geldanamycin; GR, glucocorticoid receptor; HA, hemagglutinin; Hsp90; MEF, mouse embryonic fibroblasts; Molecular chaperone; Quality control; UBR1; UPS, ubiquitin proteasome system; Ubiquitin ligase.
Figures



Similar articles
-
UBR1 promotes protein kinase quality control and sensitizes cells to Hsp90 inhibition.Exp Cell Res. 2012 Jan 1;318(1):53-60. doi: 10.1016/j.yexcr.2011.09.010. Epub 2011 Sep 29. Exp Cell Res. 2012. PMID: 21983172 Free PMC article.
-
The yeast ubr1 ubiquitin ligase participates in a prominent pathway that targets cytosolic thermosensitive mutants for degradation.G3 (Bethesda). 2012 May;2(5):619-28. doi: 10.1534/g3.111.001933. Epub 2012 May 1. G3 (Bethesda). 2012. PMID: 22670231 Free PMC article.
-
Previously unknown role for the ubiquitin ligase Ubr1 in endoplasmic reticulum-associated protein degradation.Proc Natl Acad Sci U S A. 2013 Sep 17;110(38):15271-6. doi: 10.1073/pnas.1304928110. Epub 2013 Aug 29. Proc Natl Acad Sci U S A. 2013. PMID: 23988329 Free PMC article.
-
Role of the ubiquitin-proteasome system in brain ischemia: friend or foe?Prog Neurobiol. 2014 Jan;112:50-69. doi: 10.1016/j.pneurobio.2013.10.003. Epub 2013 Oct 22. Prog Neurobiol. 2014. PMID: 24157661 Review.
-
CHIP: a co-chaperone for degradation by the proteasome.Subcell Biochem. 2015;78:219-42. doi: 10.1007/978-3-319-11731-7_11. Subcell Biochem. 2015. PMID: 25487024 Review.
Cited by
-
Bag1 Co-chaperone Promotes TRC8 E3 Ligase-dependent Degradation of Misfolded Human Ether a Go-Go-related Gene (hERG) Potassium Channels.J Biol Chem. 2017 Feb 10;292(6):2287-2300. doi: 10.1074/jbc.M116.752618. Epub 2016 Dec 20. J Biol Chem. 2017. PMID: 27998983 Free PMC article.
-
Acquired Glucocorticoid Resistance Due to Homologous Glucocorticoid Receptor Downregulation: A Modern Look at an Age-Old Problem.Cells. 2021 Sep 24;10(10):2529. doi: 10.3390/cells10102529. Cells. 2021. PMID: 34685511 Free PMC article. Review.
-
Downregulation of the Arg/N-degron Pathway Sensitizes Cancer Cells to Chemotherapy In Vivo.Mol Ther. 2020 Apr 8;28(4):1092-1104. doi: 10.1016/j.ymthe.2020.01.021. Epub 2020 Jan 21. Mol Ther. 2020. PMID: 32087767 Free PMC article.
-
UBR1 Promotes Sex-Dependent ACE2 Ubiquitination in Hypertension.Hypertension. 2025 Jan;82(1):84-95. doi: 10.1161/HYPERTENSIONAHA.124.23196. Epub 2024 Nov 27. Hypertension. 2025. PMID: 39601126
-
Ubiquitylation of nuclear receptors: new linkages and therapeutic implications.J Mol Endocrinol. 2015 Jun;54(3):R151-67. doi: 10.1530/JME-14-0308. Epub 2015 May 5. J Mol Endocrinol. 2015. PMID: 25943391 Free PMC article. Review.
References
-
- Ellis R.J. Protein misassembly: macromolecular crowding and molecular chaperones. Adv. Exp. Med. Biol. 2007;594:1–13. - PubMed
-
- Balch W.E., Morimoto R.I., Dillin A., Kelly J.W. Adapting proteostasis for disease intervention. Science. 2008;319:916–919. - PubMed
-
- Wickner S., Maurizi M.R., Gottesman S. Posttranslational quality control: folding, refolding, and degrading proteins. Science. 1999;286:1888–1893. - PubMed
-
- Hartl F.U., Bracher A., Hayer-Hartl M. Molecular chaperones in protein folding and proteostasis. Nature. 2011;475:324–332. - PubMed
-
- Hochstrasser M. Ubiquitin-dependent protein degradation. Annu. Rev. Genet. 1996;30:405–439. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

