Bacterial toxin RelE: a highly efficient ribonuclease with exquisite substrate specificity using atypical catalytic residues
- PMID: 24251350
- PMCID: PMC3910103
- DOI: 10.1021/bi401325c
Bacterial toxin RelE: a highly efficient ribonuclease with exquisite substrate specificity using atypical catalytic residues
Abstract
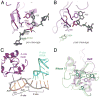
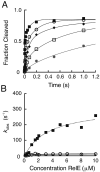
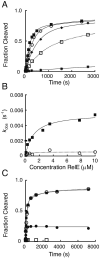
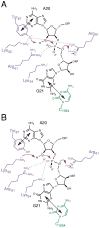
The toxin RelE is a ribosome-dependent endoribonuclease implicated in diverse cellular processes, including persistence. During amino acid starvation, RelE inhibits translation by cleaving ribosomal A-site mRNA. Although RelE is structurally similar to other microbial endoribonucleases, the active-site amino acid composition differs substantially and lacks obvious candidates for general acid-base functionality. Highly conserved RelE residues (Lys52, Lys54, Arg61, Arg81, and Tyr87) surround the mRNA scissile phosphate, and specific 16S rRNA contacts further contribute to substrate positioning. We used a single-turnover kinetic assay to evaluate the catalytic importance of individual residues in the RelE active site. Within the context of the ribosome, RelE rapidly cleaves A-site mRNA at a rate similar to those of traditional ribonucleases. Single-turnover rate constants decreased between 10(2)- and 10(6)-fold for the RelE active-site mutants of Lys52, Lys54, Arg61, and Arg81. RelE may principally promote catalysis via transition-state charge stabilization and leaving-group protonation, in addition to achieving in-line substrate positioning in cooperation with the ribosome. This kinetic analysis complements structural information to provide a foundation for understanding the molecular mechanism of this atypical endoribonuclease.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

