Growth rate-coordinated transcriptome reorganization in bacteria
- PMID: 24252326
- PMCID: PMC3840594
- DOI: 10.1186/1471-2164-14-808
Growth rate-coordinated transcriptome reorganization in bacteria
Abstract
Background: Cell growth rate reflects an organism's physiological state and largely relies on the ability of gene expression to respond to the environment. The relationship between cellular growth rate and gene expression remains unknown.
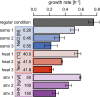
Results: Growth rate-coordinated changes in gene expression were discovered by analyzing exponentially growing Escherichia coli cells cultured under multiple defined environments, in which osmotic pressure, temperature and starvation status were varied. Gene expression analyses showed that all 3,740 genes in the genome could be simply divided into three clusters (C1, C2 and C3), which were accompanied by a generic trend in the growth rate that was coordinated with transcriptional changes. The direction of transcriptional change in C1 indicated environmental specificity, whereas those in C2 and C3 were correlated negatively and positively with growth rates, respectively. The three clusters exhibited differentiated gene functions and gene regulation task division.
Conclusions: We identified three gene clusters, exhibiting differential gene functions and distinct directions in their correlations with growth rates. Reverses in the direction of the growth rate correlated transcriptional changes and the distinguished duties of the three clusters indicated how transcriptome homeostasis is maintained to balance the total expression cost for sustaining life in new habitats.
Figures

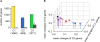



References
-
- Soupene E, Van Heeswijk WC, Plumbridge J, Stewart V, Bertenthal D, Lee H, Prasad G, Paliy O, Charernnoppakul P, Kustu S. Physiological studies of Escherichia coli strain MG1655: growth defects and apparent cross-regulation of gene expression. J Bacteriol. 2003;185(18):5611–5626. doi: 10.1128/JB.185.18.5611-5626.2003. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

