Promoter DNA methylation regulates progranulin expression and is altered in FTLD
- PMID: 24252647
- PMCID: PMC3893557
- DOI: 10.1186/2051-5960-1-16
Promoter DNA methylation regulates progranulin expression and is altered in FTLD
Abstract
Background: Frontotemporal lobar degeneration (FTLD) is a heterogeneous group of neurodegenerative diseases associated with personality changes and progressive dementia. Loss-of-function mutations in the growth factor progranulin (GRN) cause autosomal dominant FTLD, but so far the pathomechanism of sporadic FTLD is unclear.
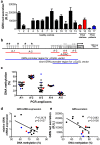
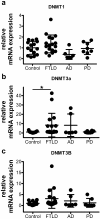
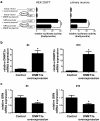
Results: We analyzed whether DNA methylation in the GRN core promoter restricts GRN expression and, thus, might promote FTLD in the absence of GRN mutations. GRN expression in human lymphoblast cell lines is negatively correlated with methylation at several CpG units within the GRN promoter. Chronic treatment with the DNA methyltransferase inhibitor 5-aza-2'-deoxycytidine (DAC) strongly induces GRN mRNA and protein levels. In a reporter assay, CpG methylation blocks transcriptional activity of the GRN core promoter. In brains of FTLD patients several CpG units in the GRN promoter are significantly hypermethylated compared to age-matched healthy controls, Alzheimer and Parkinson patients. These CpG motifs are critical for GRN promoter activity in reporter assays. Furthermore, DNA methyltransferase 3a (DNMT3a) is upregulated in FTLD patients and overexpression of DNMT3a reduces GRN promoter activity and expression.
Conclusion: These data suggest that altered DNA methylation is a novel pathomechanism for FTLD that is potentially amenable to targeted pharmacotherapy.
Figures



References
-
- Mackenzie I, Baborie A, Pickering-Brown S, Plessis D, Jaros E, Perry R, Neary D, Snowden J, Mann D. Heterogeneity of ubiquitin pathology in frontotemporal lobar degeneration: classification and relation to clinical phenotype. Acta Neuropathol. 2006;1:539–549. doi: 10.1007/s00401-006-0138-9. - DOI - PMC - PubMed
-
- Sampathu DM, Neumann M, Kwong LK, Chou TT, Micsenyi M, Truax A, Bruce J, Grossman M, Trojanowski JQ, Lee VMY. Pathological heterogeneity of frontotemporal lobar degeneration with ubiquitin-positive inclusions delineated by ubiquitin immunohistochemistry and novel monoclonal antibodies. Am J Pathol. 2006;1:1343–1352. doi: 10.2353/ajpath.2006.060438. - DOI - PMC - PubMed
-
- DeJesus-Hernandez M, Mackenzie Ian R, Boeve Bradley F, Boxer Adam L, Baker M, Rutherford Nicola J, Nicholson Alexandra M, Finch NiCole A, Flynn H, Adamson J. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron. 2011;1:245–256. doi: 10.1016/j.neuron.2011.09.011. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

