Three-dimensional electron crystallography of protein microcrystals
- PMID: 24252878
- PMCID: PMC3831942
- DOI: 10.7554/eLife.01345
Three-dimensional electron crystallography of protein microcrystals
Abstract
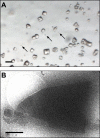
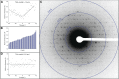
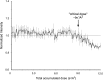
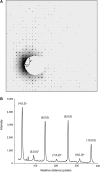
We demonstrate that it is feasible to determine high-resolution protein structures by electron crystallography of three-dimensional crystals in an electron cryo-microscope (CryoEM). Lysozyme microcrystals were frozen on an electron microscopy grid, and electron diffraction data collected to 1.7 Å resolution. We developed a data collection protocol to collect a full-tilt series in electron diffraction to atomic resolution. A single tilt series contains up to 90 individual diffraction patterns collected from a single crystal with tilt angle increment of 0.1-1° and a total accumulated electron dose less than 10 electrons per angstrom squared. We indexed the data from three crystals and used them for structure determination of lysozyme by molecular replacement followed by crystallographic refinement to 2.9 Å resolution. This proof of principle paves the way for the implementation of a new technique, which we name 'MicroED', that may have wide applicability in structural biology. DOI: http://dx.doi.org/10.7554/eLife.01345.001.
Keywords: electron cryomicroscopy (cryo-EM); electron crystallography; electron diffraction; microED; microcrystals; protein structure.
Conflict of interest statement
The authors declare that no competing interests exist.
Figures




Comment in
-
Gently does it for submicron crystals.Elife. 2013 Nov 19;2:e01662. doi: 10.7554/eLife.01662. Elife. 2013. PMID: 24252881 Free PMC article.
-
Electron crystallography goes 3D with MicroED.Nat Methods. 2014 Jan;11(1):6-7. doi: 10.1038/nmeth.2797. Nat Methods. 2014. PMID: 24524127 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

