From model specification to simulation of biologically constrained networks of spiking neurons
- PMID: 24253973
- PMCID: PMC4003408
- DOI: 10.1007/s12021-013-9208-z
From model specification to simulation of biologically constrained networks of spiking neurons
Abstract
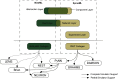
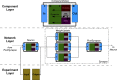
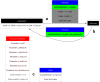
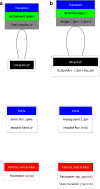
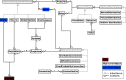
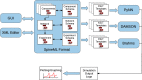
A declarative extensible markup language (SpineML) for describing the dynamics, network and experiments of large-scale spiking neural network simulations is described which builds upon the NineML standard. It utilises a level of abstraction which targets point neuron representation but addresses the limitations of existing tools by allowing arbitrary dynamics to be expressed. The use of XML promotes model sharing, is human readable and allows collaborative working. The syntax uses a high-level self explanatory format which allows straight forward code generation or translation of a model description to a native simulator format. This paper demonstrates the use of code generation in order to translate, simulate and reproduce the results of a benchmark model across a range of simulators. The flexibility of the SpineML syntax is highlighted by reproducing a pre-existing, biologically constrained model of a neural microcircuit (the striatum). The SpineML code is open source and is available at http://bimpa.group.shef.ac.uk/SpineML .
Figures
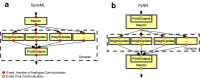
References
-
- Brette R, Rudolph M, Carnevale T, Hines M, Beeman D, Bower JM, Diesmann M, Morrison A, Goodman PH, Davison AP, Boustani SE, Destexhe A. Simulation of networks of spiking neurons: a review of tools and strategies. Journal of Computational Neuroscience. 2007;2007:349–398. doi: 10.1007/s10827-007-0038-6. - DOI - PMC - PubMed
-
- Gansner ER, Koutsofios E, North SC, Vo KP. A technique for drawing directed graphs. IEEE Transactions on Software Engineering. 1993;19(3):214–230. doi: 10.1109/32.221135. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

