Stable assembly of HIV-1 export complexes occurs cotranscriptionally
- PMID: 24255166
- PMCID: PMC3866638
- DOI: 10.1261/rna.038182.113
Stable assembly of HIV-1 export complexes occurs cotranscriptionally
Abstract
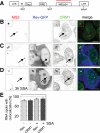
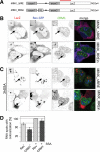
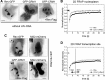
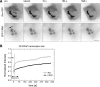
The HIV-1 Rev protein mediates export of unspliced and singly spliced viral transcripts by binding to the Rev response element (RRE) and recruiting the cellular export factor CRM1. Here, we investigated the recruitment of Rev to the transcription sites of HIV-1 reporters that splice either post- or cotranscriptionally. In both cases, we observed that Rev localized to the transcription sites of the reporters and recruited CRM1. Rev and CRM1 remained at the reporter transcription sites when cells were treated with the splicing inhibitor Spliceostatin A (SSA), showing that the proteins associate with RNA prior to or during early spliceosome assembly. Fluorescence recovery after photobleaching (FRAP) revealed that Rev and CRM1 have similar kinetics as the HIV-1 RNA, indicating that Rev, CRM1, and RRE-containing RNAs are released from the site of transcription in one single export complex. These results suggest that cotranscriptional formation of a stable export complex serves as a means to ensure efficient export of unspliced viral RNAs.
Keywords: CRM1; HIV-1 Rev; nuclear export.
Figures

References
-
- Askjaer P, Jensen TH, Nilsson J, Englmeier L, Kjems J 1998. The specificity of the CRM1-Rev nuclear export signal interaction is mediated by RanGTP. J Biol Chem 273: 33414–33422 - PubMed
-
- Bertrand E, Chartrand P, Schaefer M, Shenoy SM, Singer RH, Long RM 1998. Localization of ASH1 mRNA particles in living yeast. Mol Cell 2: 437–445 - PubMed
-
- Chang DD, Sharp PA 1989. Regulation by HIV Rev depends upon recognition of splice sites. Cell 59: 789–795 - PubMed
-
- Cullen BR 1998. Retroviruses as model systems for the study of nuclear RNA export pathways. Virology 249: 203–210 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
