Identification of new p53 target microRNAs by bioinformatics and functional analysis
- PMID: 24256616
- PMCID: PMC4225545
- DOI: 10.1186/1471-2407-13-552
Identification of new p53 target microRNAs by bioinformatics and functional analysis
Abstract
Background: The tumor suppressor p53 is a sequence-specific transcription factor that regulates an extensive network of coding genes, long non-coding RNAs and microRNAs, that establish intricate gene regulatory circuits influencing many cellular responses beyond the prototypical control of cell cycle, apoptosis and DNA repair.
Methods: Using bioinformatic approaches, we identified an additional group of candidate microRNAs (miRs) under direct p53 transcriptional control. To validate p53 family-mediated responsiveness of the newly predicted target miRs we first evaluated the potential for wild type p53, p63β and p73β to transactivate from p53 response elements (REs) mapped in the miR promoters, using an established yeast-based assay.
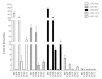
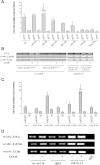
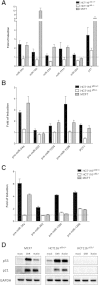
Results: The REs found in miR-10b, -23b, -106a, -151a, -191, -198, -202, -221, -320, -1204, -1206 promoters were responsive to p53 and 8 of them were also responsive to p63β or p73β. The potential for germline p53 mutations to drive transactivation at selected miR-associated REs was also examined. Chromatin Immuno-Precipitation (ChIP) assays conducted in doxorubicin-treated MCF7 cells and HCT116 p53+/+ revealed moderate induction of p53 occupancy at the miR-202, -1204, -1206, -10b RE-containing sites, while weak occupancy was observed for the miR-23b-associated RE only in MCF7 cells. RT-qPCR analyses cells showed modest doxorubicin- and/or Nutlin-dependent induction of the levels of mature miR-10b, -23b, -151a in HCT116 p53+/+ and MCF7 cells. The long noncoding RNA PVT1 comprising miR-1204 and -1206 was weakly induced only in HCT116 p53+/+ cells, but the mature miRs were not detected. miR-202 expression was not influenced by p53-activating stimuli in our cell systems.
Conclusions: Our study reveals additional miRs, particularly miR-10b and miR-151a, that could be directly regulated by the p53-family of transcription factors and contribute to the tuning of p53-induced responses.
Figures

Similar articles
-
The ferredoxin reductase gene is regulated by the p53 family and sensitizes cells to oxidative stress-induced apoptosis.Oncogene. 2002 Oct 17;21(47):7195-204. doi: 10.1038/sj.onc.1205862. Oncogene. 2002. PMID: 12370809
-
p73 function is inhibited by tumor-derived p53 mutants in mammalian cells.Mol Cell Biol. 1999 Feb;19(2):1438-49. doi: 10.1128/MCB.19.2.1438. Mol Cell Biol. 1999. PMID: 9891077 Free PMC article.
-
Induction of the small heat shock protein alphaB-crystallin by genotoxic stress is mediated by p53 and p73.Breast Cancer Res Treat. 2010 Jul;122(1):159-68. doi: 10.1007/s10549-009-0542-7. Epub 2009 Sep 24. Breast Cancer Res Treat. 2010. PMID: 19777343 Free PMC article.
-
The guardians of the genome (p53, TA-p73, and TA-p63) are regulators of tumor suppressor miRNAs network.Cancer Metastasis Rev. 2010 Dec;29(4):613-39. doi: 10.1007/s10555-010-9257-9. Cancer Metastasis Rev. 2010. PMID: 20922462 Review.
-
The microRNA feedback regulation of p63 in cancer progression.Oncotarget. 2015 Apr 20;6(11):8434-53. doi: 10.18632/oncotarget.3020. Oncotarget. 2015. PMID: 25726529 Free PMC article. Review.
Cited by
-
HDAC5-mediated exosomal Maspin and miR-151a-3p as biomarkers for enhancing radiation treatment sensitivity in hepatocellular carcinoma.Biomater Res. 2023 Dec 15;27(1):134. doi: 10.1186/s40824-023-00467-7. Biomater Res. 2023. PMID: 38102691 Free PMC article.
-
Downregulated miR-23b-3p expression acts as a predictor of hepatocellular carcinoma progression: A study based on public data and RT-qPCR verification.Int J Mol Med. 2018 May;41(5):2813-2831. doi: 10.3892/ijmm.2018.3513. Epub 2018 Feb 23. Int J Mol Med. 2018. PMID: 29484429 Free PMC article.
-
Regulation mechanism and pathogenic role of lncRNA plasmacytoma variant translocation 1 (PVT1) in human diseases.Genes Dis. 2022 Jun 29;10(3):901-914. doi: 10.1016/j.gendis.2022.05.037. eCollection 2023 May. Genes Dis. 2022. PMID: 37396533 Free PMC article. Review.
-
Influence of pentoxifylline on gene expression of PAG1/ miR-1206/ SNHG14 in ischemic heart disease.Biochem Biophys Rep. 2021 Jan 27;25:100911. doi: 10.1016/j.bbrep.2021.100911. eCollection 2021 Mar. Biochem Biophys Rep. 2021. PMID: 33553684 Free PMC article.
-
The 5'-untranslated region of p16INK4a melanoma tumor suppressor acts as a cellular IRES, controlling mRNA translation under hypoxia through YBX1 binding.Oncotarget. 2015 Nov 24;6(37):39980-94. doi: 10.18632/oncotarget.5387. Oncotarget. 2015. PMID: 26498684 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

