To tree or not to tree? Genome-wide quantification of recombination and reticulate evolution during the diversification of strict intracellular bacteria
- PMID: 24259310
- PMCID: PMC3879967
- DOI: 10.1093/gbe/evt178
To tree or not to tree? Genome-wide quantification of recombination and reticulate evolution during the diversification of strict intracellular bacteria
Abstract
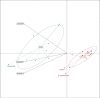
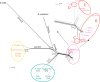
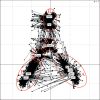
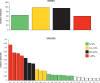
It is well known that horizontal gene transfer (HGT) is a major force in the evolution of prokaryotes. During the adaptation of a bacterial population to a new ecological niche, and particularly for intracellular bacteria, selective pressures are shifted and ecological niches reduced, resulting in a lower rate of genetic connectivity. HGT and positive selection are therefore two important evolutionary forces in microbial pathogens that drive adaptation to new hosts. In this study, we use genomic distance analyses, phylogenomic networks, tree topology comparisons, and Bayesian inference methods to investigate to what extent HGT has occurred during the evolution of the genus Rickettsia, the effect of the use of different genomic regions in estimating reticulate evolution and HGT events, and the link of these to host range. We show that ecological specialization restricts recombination occurrence in Rickettsia, but other evolutionary processes and genome architecture are also important for the occurrence of HGT. We found that recombination, genomic rearrangements, and genome conservation all show evidence of network-like evolution at whole-genome scale. We show that reticulation occurred mainly, but not only, during the early Rickettsia radiation, and that core proteome genes of every major functional category have experienced reticulated evolution and possibly HGT. Overall, the evolution of Rickettsia bacteria has been tree-like, with evidence of HGT and reticulated evolution for around 10-25% of the core Rickettsia genome. We present evidence of extensive recombination/incomplete lineage sorting (ILS) during the radiation of the genus, probably linked with the emergence of intracellularity in a wide range of hosts.
Keywords: Rickettsia; bacterial speciation; horizontal gene transfer; phylogenomic networks; reticulate evolution.
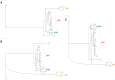
Figures

Similar articles
-
Algorithms for computing parsimonious evolutionary scenarios for genome evolution, the last universal common ancestor and dominance of horizontal gene transfer in the evolution of prokaryotes.BMC Evol Biol. 2003 Jan 6;3:2. doi: 10.1186/1471-2148-3-2. Epub 2003 Jan 6. BMC Evol Biol. 2003. PMID: 12515582 Free PMC article.
-
The Phylogeny of Rickettsia Using Different Evolutionary Signatures: How Tree-Like is Bacterial Evolution?Syst Biol. 2016 Mar;65(2):265-79. doi: 10.1093/sysbio/syv084. Epub 2015 Nov 11. Syst Biol. 2016. PMID: 26559010 Free PMC article.
-
Phylogenomic species tree estimation in the presence of incomplete lineage sorting and horizontal gene transfer.BMC Genomics. 2015;16 Suppl 10(Suppl 10):S1. doi: 10.1186/1471-2164-16-S10-S1. Epub 2015 Oct 2. BMC Genomics. 2015. PMID: 26450506 Free PMC article.
-
What Is Speciation?PLoS Genet. 2016 Mar 31;12(3):e1005860. doi: 10.1371/journal.pgen.1005860. eCollection 2016 Mar. PLoS Genet. 2016. PMID: 27030977 Free PMC article. Review.
-
Phylogenomic networks.Trends Microbiol. 2011 Oct;19(10):483-91. doi: 10.1016/j.tim.2011.07.001. Epub 2011 Aug 3. Trends Microbiol. 2011. PMID: 21820313 Review.
Cited by
-
Reticulate evolution is favored in influenza niche switching.Proc Natl Acad Sci U S A. 2016 May 10;113(19):5335-9. doi: 10.1073/pnas.1522921113. Epub 2016 Apr 25. Proc Natl Acad Sci U S A. 2016. PMID: 27114508 Free PMC article.
-
Transcriptional Analysis of the Conjugal Transfer Genes of Rickettsia bellii RML 369-C.PLoS One. 2015 Sep 9;10(9):e0137214. doi: 10.1371/journal.pone.0137214. eCollection 2015. PLoS One. 2015. PMID: 26352829 Free PMC article.
-
Contributions of ancestral inter-species recombination to the genetic diversity of extant Streptomyces lineages.ISME J. 2016 Jul;10(7):1731-41. doi: 10.1038/ismej.2015.230. Epub 2016 Feb 5. ISME J. 2016. PMID: 26849310 Free PMC article.
-
A Robust ANOVA Approach to Estimating a Phylogeny from Multiple Genes.Mol Biol Evol. 2015 Aug;32(8):2186-94. doi: 10.1093/molbev/msv084. Epub 2015 Apr 3. Mol Biol Evol. 2015. PMID: 25841490 Free PMC article.
-
Reevaluation of Parasynechococcus-like Strains and Genomic Analysis of Their Microsatellites and Compound Microsatellites.Plants (Basel). 2022 Apr 13;11(8):1060. doi: 10.3390/plants11081060. Plants (Basel). 2022. PMID: 35448788 Free PMC article.
References
-
- Amiri H, Davids W, Andersson SGE. Birth and death of orphan genes in Rickettsia. Mol Biol Evol. 2003;20:1575–1587. - PubMed
-
- Andersson JO, Andersson SGE. Genome degradation is an ongoing process in Rickettsia. Mol Biol Evol. 1999;16:1178–1191. - PubMed
-
- Ané C, Larget B, Baum DA, Smith SD, Rokas A. Bayesian estimation of concordance among gene trees. Mol Biol Evol. 2007;24:412–426. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

