Argonaute 2 sustains the gene expression program driving human monocytic differentiation of acute myeloid leukemia cells
- PMID: 24263100
- PMCID: PMC3847328
- DOI: 10.1038/cddis.2013.452
Argonaute 2 sustains the gene expression program driving human monocytic differentiation of acute myeloid leukemia cells
Abstract
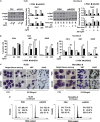
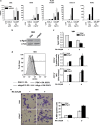
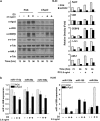
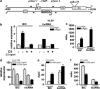
MicroRNAs are key regulators of many biological processes, including cell differentiation. These small RNAs exert their function assembled in the RNA-induced silencing complexes (RISCs), where members of Argonaute (Ago) family of proteins provide a unique platform for target recognition and gene silencing. Here, by using myeloid cell lines and primary blasts, we show that Ago2 has a key role in human monocytic cell fate determination and in LPS-induced inflammatory response of 1,25-dihydroxyvitamin D3 (D3)-treated myeloid cells. The silencing of Ago2 impairs the D3-dependent miR-17-5p/20a/106a, miR-125b and miR-155 downregulation, the accumulation of their translational targets AML1, VDR and C/EBPβ and monocytic cell differentiation. Moreover, we show that Ago2 is recruited on miR-155 host gene promoter and on the upstream region of an overlapping antisense lncRNA, determining their epigenetic silencing, and miR-155 downregulation. These findings highlight Ago2 as a new factor in myeloid cell fate determination in acute myeloid leukemia cells.
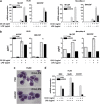
Figures

Similar articles
-
1,25-Dihydroxyvitamin D3 induces monocytic differentiation of human myeloid leukemia cells by regulating C/EBPβ expression through MEF2C.J Steroid Biochem Mol Biol. 2015 Apr;148:132-7. doi: 10.1016/j.jsbmb.2014.11.016. Epub 2014 Nov 20. J Steroid Biochem Mol Biol. 2015. PMID: 25448741 Free PMC article.
-
MicroRNA-29a and microRNA-142-3p are regulators of myeloid differentiation and acute myeloid leukemia.Blood. 2012 May 24;119(21):4992-5004. doi: 10.1182/blood-2011-10-385716. Epub 2012 Apr 4. Blood. 2012. PMID: 22493297
-
Regulation of Expression of CEBP Genes by Variably Expressed Vitamin D Receptor and Retinoic Acid Receptor α in Human Acute Myeloid Leukemia Cell Lines.Int J Mol Sci. 2018 Jun 29;19(7):1918. doi: 10.3390/ijms19071918. Int J Mol Sci. 2018. PMID: 29966306 Free PMC article.
-
Regulation of the MIR155 host gene in physiological and pathological processes.Gene. 2013 Dec 10;532(1):1-12. doi: 10.1016/j.gene.2012.12.009. Epub 2012 Dec 14. Gene. 2013. PMID: 23246696 Review.
-
C/EBPα in normal and malignant myelopoiesis.Int J Hematol. 2015 Apr;101(4):330-41. doi: 10.1007/s12185-015-1764-6. Epub 2015 Mar 10. Int J Hematol. 2015. PMID: 25753223 Free PMC article. Review.
Cited by
-
Analysis of long non-coding RNA expression profiles in clear cell renal cell carcinoma.Oncol Lett. 2017 Sep;14(3):2757-2764. doi: 10.3892/ol.2017.6563. Epub 2017 Jul 8. Oncol Lett. 2017. PMID: 28928816 Free PMC article.
-
Argonaute 2 Restores Erectile Function by Enhancing Angiogenesis and Reducing Reactive Oxygen Species Production in Streptozotocin (STZ)-Induced Type-1 Diabetic Mice.Int J Mol Sci. 2023 Feb 2;24(3):2935. doi: 10.3390/ijms24032935. Int J Mol Sci. 2023. PMID: 36769259 Free PMC article.
-
The varied roles of nuclear argonaute-small RNA complexes and avenues for therapy.Mol Ther Nucleic Acids. 2014 Oct 14;3(10):e203. doi: 10.1038/mtna.2014.54. Mol Ther Nucleic Acids. 2014. PMID: 25313622 Free PMC article. Review.
-
Thymic Epithelial Tumors phenotype relies on miR-145-5p epigenetic regulation.Mol Cancer. 2017 May 10;16(1):88. doi: 10.1186/s12943-017-0655-2. Mol Cancer. 2017. PMID: 28486946 Free PMC article.
-
The Potential of Vitamin D-Regulated Intracellular Signaling Pathways as Targets for Myeloid Leukemia Therapy.J Clin Med. 2015 Mar 25;4(4):504-34. doi: 10.3390/jcm4040504. J Clin Med. 2015. PMID: 26239344 Free PMC article. Review.
References
-
- Rosenbauer F, Tenen DG. Transcription factors in myeloid development: balancing differentiation with transformation. Nat Rev Immunol. 2007;7:105–117. - PubMed
-
- Fazi F, Nervi C. MicroRNA: basic mechanisms and transcriptional regulatory networks for cell fate determination. Cardiovasc Res. 2008;79:553–561. - PubMed
-
- García P, Frampton J. Hematopoietic lineage commitment: miRNAs add specificity to a widely expressed transcription factor. Developmental cell. 2008;14:815–816. - PubMed
-
- Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR, et al. Proposed revised criteria for the classification of acute myeloid leukemia. A report of the French-American-British Cooperative Group. Ann Intern Med. 1985;103:620–625. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

