Tsix RNA and the germline factor, PRDM14, link X reactivation and stem cell reprogramming
- PMID: 24268575
- PMCID: PMC3950835
- DOI: 10.1016/j.molcel.2013.10.023
Tsix RNA and the germline factor, PRDM14, link X reactivation and stem cell reprogramming
Abstract
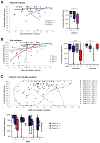
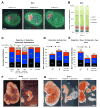
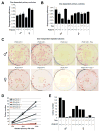
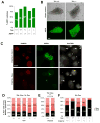
Transitions between pluripotent and differentiated states are marked by dramatic epigenetic changes. Cellular differentiation is tightly linked to X chromosome inactivation (XCI), whereas reprogramming to induced pluripotent stem cells (iPSCs) is associated with X chromosome reactivation (XCR). XCR reverses the silent state of the inactive X, occurring in mouse blastocysts and germ cells. In spite of its importance, little is known about underlying mechanisms. Here, we examine the role of the long noncoding Tsix RNA and the germline factor, PRDM14. In blastocysts, XCR is perturbed by mutation of either Tsix or Prdm14. In iPSCs, XCR is disrupted only by PRDM14 deficiency, which also affects iPSC derivation and maintenance. We show that Tsix and PRDM14 directly link XCR to pluripotency: first, PRDM14 represses Rnf12 by recruiting polycomb repressive complex 2; second, Tsix enables PRDM14 to bind Xist. Thus, our study provides functional and mechanistic links between cellular and X chromosome reprogramming.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures


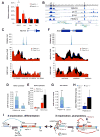
Similar articles
-
Coupling of X-chromosome reactivation with the pluripotent stem cell state.RNA Biol. 2014;11(7):798-807. doi: 10.4161/rna.29779. Epub 2014 Aug 19. RNA Biol. 2014. PMID: 25137047 Free PMC article.
-
Histone demethylation maintains Prdm14 and Tsix expression and represses xIst in embryonic stem cells.PLoS One. 2015 May 20;10(5):e0125626. doi: 10.1371/journal.pone.0125626. eCollection 2015. PLoS One. 2015. PMID: 25993097 Free PMC article.
-
A PRC2-dependent repressive role of PRDM14 in human embryonic stem cells and induced pluripotent stem cell reprogramming.Stem Cells. 2013 Apr;31(4):682-92. doi: 10.1002/stem.1307. Stem Cells. 2013. PMID: 23280602
-
New Insights into X-Chromosome Reactivation during Reprogramming to Pluripotency.Cells. 2020 Dec 17;9(12):2706. doi: 10.3390/cells9122706. Cells. 2020. PMID: 33348832 Free PMC article. Review.
-
Reactivation of the inactive X chromosome in development and reprogramming.Cell Mol Life Sci. 2013 Jul;70(14):2443-61. doi: 10.1007/s00018-012-1174-3. Epub 2012 Sep 30. Cell Mol Life Sci. 2013. PMID: 23052214 Free PMC article. Review.
Cited by
-
Dysregulation of long non-coding RNA in breast cancer: an overview of mechanism and clinical implication.Oncotarget. 2017 Jan 17;8(3):5508-5522. doi: 10.18632/oncotarget.12537. Oncotarget. 2017. PMID: 27732939 Free PMC article. Review.
-
Orchestrating Asymmetric Expression: Mechanisms behind Xist Regulation.Epigenomes. 2024 Feb 1;8(1):6. doi: 10.3390/epigenomes8010006. Epigenomes. 2024. PMID: 38390897 Free PMC article. Review.
-
Chromosome compartments on the inactive X guide TAD formation independently of transcription during X-reactivation.Nat Commun. 2021 Jun 9;12(1):3499. doi: 10.1038/s41467-021-23610-1. Nat Commun. 2021. PMID: 34108480 Free PMC article.
-
Sexual Dimorphism of the Heart: Genetics, Epigenetics, and Development.Front Cardiovasc Med. 2021 May 26;8:668252. doi: 10.3389/fcvm.2021.668252. eCollection 2021. Front Cardiovasc Med. 2021. PMID: 34124200 Free PMC article. Review.
-
Enhanced chromatin accessibility contributes to X chromosome dosage compensation in mammals.Genome Biol. 2021 Nov 1;22(1):302. doi: 10.1186/s13059-021-02518-5. Genome Biol. 2021. PMID: 34724962 Free PMC article.
References
-
- Barakat TS, Gribnau J. X chromosome inactivation in the cycle of life. Development. 2012;139:2085–2089. - PubMed
-
- Brown CJ, Hendrich BD, Rupert JL, Lafreniere RG, Xing Y, Lawrence J, Willard HF. The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell. 1992;71:527–542. - PubMed
-
- Chan YS, Goke J, Lu X, Venkatesan N, Feng B, Su IH, Ng HH. A PRC2-dependent repressive role of PRDM14 in human embryonic stem cells and induced pluripotent stem cell reprogramming. Stem Cells. 2012;31:682–692. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

