Quantitative proteomic analysis in HCV-induced HCC reveals sets of proteins with potential significance for racial disparity
- PMID: 24283668
- PMCID: PMC3850534
- DOI: 10.1186/1479-5876-11-239
Quantitative proteomic analysis in HCV-induced HCC reveals sets of proteins with potential significance for racial disparity
Abstract
Background: The incidence and mortality of hepatitis C virus (HCV)-induced hepatocellular carcinoma (HCC) is higher in African Americans (AA) than other racial/ethnic groups in the U.S., but the reasons for this disparity are unknown. There is an urgent need for the discovery of novel molecular signatures for HCV disease progression to understand the underlying biological basis for this cancer rate disparity to improve the clinical outcome.
Methods: We performed differential proteomics with isobaric labeling tags for relative and absolute quantitation (iTRAQ) and MS/MS analysis to identify proteins differentially expressed in cirrhotic (CIR) and HCC as compared to normal tissues of Caucasian American (CA) patients. The raw data were analyzed using the ProteinPilot v3.0. Searches were performed against all known sequences populating the Swiss-Prot, Refseq, and TrEMBL databases. Quality control analyses were accomplished using pairwise correlation plots, boxplots, principal component analysis, and unsupervised hierarchical clustering. Supervised analysis was carried out to identify differentially expressed proteins. Candidates were validated in independent cohorts of CA and AA tissues by qRT-PCR or Western blotting.
Results: A total of 238 unique proteins were identified. Of those, around 15% were differentially expressed between normal, CIR & HCC groups. Target validation demonstrates racially distinct alteration in the expression of certain proteins. For example, the mRNA expression levels of transferrin (TF) were 2 and18-fold higher in CIR and HCC in AA as compared to CA. Similarly; the expression of Apolipoprotein A1 (APOA1) was 7-fold higher in HCC of AA. This increase was mirrored in the protein expression levels. Interestingly, the level of hepatocyte nuclear factor4a (HNF4a) protein was down regulated in AA, whereas repression of transcription is seen more in CA compared to AA. These data suggest that racial disparities in HCC could be a consequence of differential dysregulation of HNF4a transcriptional activity.
Conclusion: This study identifies novel molecular signatures in HCV-induced HCC using iTRAQ-based tissue proteomics. The proteins identified will further enhance a molecular explanation to the biochemical mechanism(s) that may play a role in HCC racial disparities.
Figures



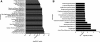


Similar articles
-
Differential Expression of MicroRNAs in Hepatitis C Virus-Mediated Liver Disease Between African Americans and Caucasians: Implications for Racial Health Disparities.Gene Expr. 2017 Feb 10;17(2):89-98. doi: 10.3727/105221616X693594. Epub 2016 Oct 19. Gene Expr. 2017. PMID: 27765085 Free PMC article.
-
Proteome Differences between Hepatitis B Virus Genotype-B- and Genotype-C-Induced Hepatocellular Carcinoma Revealed by iTRAQ-Based Quantitative Proteomics.J Proteome Res. 2016 Feb 5;15(2):487-98. doi: 10.1021/acs.jproteome.5b00838. Epub 2016 Jan 14. J Proteome Res. 2016. PMID: 26709725
-
Genes involved in viral carcinogenesis and tumor initiation in hepatitis C virus-induced hepatocellular carcinoma.Mol Med. 2009 Mar-Apr;15(3-4):85-94. doi: 10.2119/molmed.2008.00110. Epub 2008 Dec 15. Mol Med. 2009. PMID: 19098997 Free PMC article.
-
Oncogenic long noncoding RNA MALAT1 and HCV-related hepatocellular carcinoma.Biomed Pharmacother. 2018 Jun;102:653-669. doi: 10.1016/j.biopha.2018.03.105. Epub 2018 Apr 5. Biomed Pharmacother. 2018. PMID: 29604585
-
Current progress in proteomic study of hepatitis C virus-related human hepatocellular carcinoma.Expert Rev Proteomics. 2005 Aug;2(4):589-601. doi: 10.1586/14789450.2.4.589. Expert Rev Proteomics. 2005. PMID: 16097891 Review.
Cited by
-
Proteomic Upregulation of Fatty Acid Synthase and Fatty Acid Binding Protein 5 and Identification of Cancer- and Race-Specific Pathway Associations in Human Prostate Cancer Tissues.J Cancer. 2016 Jul 5;7(11):1452-64. doi: 10.7150/jca.15860. eCollection 2016. J Cancer. 2016. PMID: 27471561 Free PMC article.
-
Features of hepatocellular carcinoma in Hispanics differ from African Americans and non-Hispanic Whites.World J Hepatol. 2017 Mar 8;9(7):391-400. doi: 10.4254/wjh.v9.i7.391. World J Hepatol. 2017. PMID: 28321275 Free PMC article.
-
Study of Cathepsin B inhibition in VEGFR TKI treated human renal cell carcinoma xenografts.Oncogenesis. 2019 Feb 22;8(3):15. doi: 10.1038/s41389-019-0121-7. Oncogenesis. 2019. PMID: 30796200 Free PMC article.
-
DNA methyltransferase 1, 3a, and 3b expression in hepatitis C associated human hepatocellular carcinoma and their clinicopathological association.Tumour Biol. 2016 Aug;37(8):10487-97. doi: 10.1007/s13277-016-4941-1. Epub 2016 Feb 5. Tumour Biol. 2016. PMID: 26850594
-
Racial/ethnic disparities in hepatocellular carcinoma treatment and survival in California, 1988-2012.World J Gastroenterol. 2016 Oct 14;22(38):8584-8595. doi: 10.3748/wjg.v22.i38.8584. World J Gastroenterol. 2016. PMID: 27784971 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

