The innate growth bistability and fitness landscapes of antibiotic-resistant bacteria
- PMID: 24288338
- PMCID: PMC4059556
- DOI: 10.1126/science.1237435
The innate growth bistability and fitness landscapes of antibiotic-resistant bacteria
Abstract
To predict the emergence of antibiotic resistance, quantitative relations must be established between the fitness of drug-resistant organisms and the molecular mechanisms conferring resistance. These relations are often unknown and may depend on the state of bacterial growth. To bridge this gap, we have investigated Escherichia coli strains expressing resistance to translation-inhibiting antibiotics. We show that resistance expression and drug inhibition are linked in a positive feedback loop arising from an innate, global effect of drug-inhibited growth on gene expression. A quantitative model of bacterial growth based on this innate feedback accurately predicts the rich phenomena observed: a plateau-shaped fitness landscape, with an abrupt drop in the growth rates of cultures at a threshold drug concentration, and the coexistence of growing and nongrowing populations, that is, growth bistability, below the threshold.
Figures



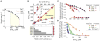

References
-
- Alanis AJ. Resistance to antibiotics: are we in the post-antibiotic era? Archives of medical research. 2005;36:697–705. - PubMed
-
- World Health Organization. The evolving threat of antimicrobial resistance: Options for action. World Health Organization; 2012.
-
- Martínez JL, Baquero F, Andersson DI. Predicting antibiotic resistance. Nat Rev Microbiol. 2007;5:958–65. - PubMed
-
- MacLean RC, Hall AR, Perron GG, Buckling A. The population genetics of antibiotic resistance: integrating molecular mechanisms and treatment contexts. Nat Rev Genet. 2010;11:405–14. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

