Disruption of autoregulatory feedback by a mutation in a remote, ultraconserved PAX6 enhancer causes aniridia
- PMID: 24290376
- PMCID: PMC3852925
- DOI: 10.1016/j.ajhg.2013.10.028
Disruption of autoregulatory feedback by a mutation in a remote, ultraconserved PAX6 enhancer causes aniridia
Abstract
The strictly regulated expression of most pleiotropic developmental control genes is critically dependent on the activity of long-range cis-regulatory elements. This was revealed by the identification of individuals with a genetic condition lacking coding-region mutations in the gene commonly associated with the disease but having a variety of nearby chromosomal abnormalities, collectively described as cis-ruption disease cases. The congenital eye malformation aniridia is caused by haploinsufficiency of the developmental regulator PAX6. We discovered a de novo point mutation in an ultraconserved cis-element located 150 kb downstream from PAX6 in an affected individual with intact coding region and chromosomal locus. The element SIMO acts as a strong enhancer in developing ocular structures. The mutation disrupts an autoregulatory PAX6 binding site, causing loss of enhancer activity, resulting in defective maintenance of PAX6 expression. These findings reveal a distinct regulatory mechanism for genetic disease by disruption of an autoregulatory feedback loop critical for maintenance of gene expression through development.
Copyright © 2013 The American Society of Human Genetics. Published by Elsevier Inc. All rights reserved.
Figures



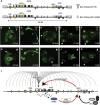
References
-
- Kleinjan D.J., Coutinho P. Cis-ruption mechanisms: disruption of cis-regulatory control as a cause of human genetic disease. Brief. Funct. Genomics Proteomics. 2009;8:317–332. - PubMed
-
- Fantes J., Redeker B., Breen M., Boyle S., Brown J., Fletcher J., Jones S., Bickmore W., Fukushima Y., Mannens M. Aniridia-associated cytogenetic rearrangements suggest that a position effect may cause the mutant phenotype. Hum. Mol. Genet. 1995;4:415–422. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

